Screening of Bacterial Isolates from Seafood-Wastes for Chitin Degrading Enzyme Activity
- 1. Amity Institute of Biotechnology, Amity University, India
- 2. PDDYPatil University, Navi Mumbai, Maharashtra, India
- 3. Biomedical Sciences Research Institute, Ulster University, UK
ABSTARCT
The safer removal of chitin is important for controlling accumulation of chitin in environment and agriculture. Chitin a long-chain polymer is a glucosederivative and a primary component of fungal cell-wall, the exoskeleton of molluscs, fish, and insects. Chitinase enzyme (E.C. 3.2.1.14) is able to digest chitin, hydrolyzing the 1-4 linkages between the N-acetyleglucosamine molecules. Present work was undertaken for the isolation and characterisation of chitinolytic bacteria from waste-seafood. Seventy-one indigenous strains were isolated from a variety of marine-sources, including scales of sea-fish, shrimps, prawns and crabs; of which 4isolates were identified as Alcaligenes spp., 2as Kurthiaspp., 1 as Acinetobactor and 1 as Bacillus spp. These were higher chitinase producers with enzyme units ranging from 12.5 to 35.41 micromoles/min/ml. These chitinase producers may prove to be industrially beneficial for their enzyme production for degradation of chitin in environment clean-up, polluted with decaying sea-food and wastes originated from their processing.
CITATION
Thomas S, Patil AB, Salgaonkar PN, Shrivastava S, Nigam PS (2020) Screening of Bacterial Isolates from Seafood-Wastes for Chitin Degrading Enzyme Activity. Chem Eng Process Tech 5(1): 1059.
INTRODUCTION
Chitin, a β-1,4-linked polymer of N-acetyl-β-D-glucosamine (NAG), is the second most abundant polysaccharide in nature after cellulose. Chitin is produced by many aquatic and terrestrial organisms in different forms [1]. It is a major cell wall constituent of higher fungi (chitridiomycetes, ascomycetes, basidiomycetes and deuteromycetes), insect exoskeletons and crustacean shells [2]. About 1011 tonnes of chitin is produced annually in the aquatic biosphere alone [3,4], arising from processing of seafood - crab, shrimp and krill shells [5]. Its complex structure chitin is resistance to natural process of degradation, creating disposal problems in environment due to its presence in sea-food wastes. Second major issue is experienced in agriculture industry, where chitin in pests is a protective-barrier against action of pesticides [6]. Despite being a potential renewable resource, most of the chitinous wastes are disposed through ocean dumping, incineration and land filling; owing to lack of cheap and commercially feasible methods for its processing leading to wastage of natural resource, economic loss and environmental pollution [7]. In India alone 60-80,000 tonnes of chitinous wastes are generated, making it necessary to design alternative methods for its disposal and utilization [8]. Chitin and chitosan nanofibers could be extracted from crustacean wastes for the development of products formedicine, and tissue engineering [9].
Chitin is insoluble in water, dilute and concentrated alkalis, alcohol and other organic solvents. These major issues have led to increased interest in chitin?hydrolyzing enzymes: chitinases. (E.C. 3.2.1.14) hydrolyzing the 1-4 linkages between the NAG molecules [10]. These glycosyl hydrolases (GHs) are about 20- 90 kDa in size and are present in various organisms [11,12]. In chitin-containing organisms, chitinases play an important role in normal life cycle functions such as morphogenesis and cell division, whereas plants produce chitinases as part of their defence against fungal pathogens. Chitinases derived from a variety of prokaryotic and eukaryotic organisms constitute the family 18of GHs, whereas those derived from only higher plants and Gram-positive bacterium Streptomyces constitute the family-19. Family-20 includes N-acetylglucosaminidase from Vibrio harveyi and N-acetylhexosaminidase from Dictyostelium discoideum and human [13]. Endo-chitinases cleave randomly in the chitin chain and exo-chitinasescleave off chitobiose or chitotriose from the reducing or the non-reducing ends of the chitin chain. Microbial degradation of chitin not only leads to recycling of nutrients in the environment but also facilitates the treatment of chitinous wastes, mycolytic enzyme preparation along with generation of useful products viz. chitooligosaccharides, N-acetylglucosamine and single cell proteins [14-16]. Chitinases have found major applications in the field of agriculture, medicine, biotechnology, food technology, waste management and industry. They act as potential biocontrol agents to fungal phytopathogens [10], mosquitoes and harmful insects [17] as well as aidin the development of fungal resistant plant/crops that carry a chitinase transgene [18-20]. Detection of chitinolytic microorganisms from natural sources is vital in the isolation of chitinase producing strains. The aim of this study was to isolate the most prominent chitinolytic microorganisms from different marine wastes and water effluent as well as analyze the hydrolysis products formed as a result of the fermentation by these isolates.
These enzymes break down the 1→4 β?glycoside bond of N? acetyl d?glucosamine in chitin to produce mono? and oligomers. The inducible nature of chitinases, low activity of synthesized enzymes, and inertia of the substrate are only a few of the problems that need to be solved by biotechnology. Our project aims to screen bacteria capable of growing in Chitin containing medium and selection of strains which are effective producer of chitinolytic enzymes, with objective to promote their use for environment and in agriculutre.
MATERIALS AND METHODS
Sample preparation for isolation of chitinolyticbacteria
Chitinase producing bacterial strains were isolated from marine sources. Several samples were collected from different regions (Alibag, Vashi and Belapur of Mumbai city) in west coastal state of India. Sources for samples included wastewater effluent from an industry in Taloja, white prawn shells (Fenneropenaeusindicus, formerly Penaeusindicus), Tiger-prawn shells (Penaeus monodon), sea-fish scales and sea crab shells (Charybdis cruciata). Each sample was separately crushed in a sterilized mortar-pestle under aseptic conditions using sterile water. Filtrates collectedwere serially diluted in sterile saline solution, 200 µl of 10-8, 10-12 and 10-15 dilution was inoculatedon nutrient agar plates and incubated at 37°C for 24 hours. Colonies obtained from the master plates were isolated and taken for further studies.
Cells from purified colonies of Isolates were spot-inoculated on sterile minimal M9 medium [21], with minor modification containing colloidal chitin pH 6.5 [18] and incubated at 37°C for 24-48 hours. Isolates were maintained by sub-culturing after every two weeks. These were screened for secretion of chitinase on containing chitin medium.
Identification of chitinolytic microorganisms
Isolates screened positive for chitinase production were tested using Gram’s staining method [22], and identified through 16S rDNA sequencing carried out at the sequencing facility of Microbial Culture Collection at National Centre for Cell Sciences (NCCS), Pune, India. Sequences of 16S rDNA obtained were analyzed for the identification and phylogenetic relationships were studied using http://blast.ncbi.nlm.nih.gov/Blast.cgi; www. cme.msu.edu/RDP/html/index.html and www.ebi.ac.uk/Tools/ msa/clustalw2/.
Chitinase Production
Microorganisms screened positive for chitinase production were selected for making starter inoculum culture by inoculating loop-full of colonies in 25 ml of M9 minimal medium containing 0.5% chitin flakes (pH 7) and incubated at 37°C at 100rpm for 24 h. Minimal medium containing chitin as the sole carbon source was used as enzyme production mediumin fermentation process. Medium consisted of (g L-1): K2 HPO4 , 0.7; KH2 PO4 , 0.3; sodium citrate, 0.05; MgSO4 .7H2 O, 0.01; ammonium sulphate, 0.1; acid swollen chitin, 1% (w/v) [23] and colloidal chitin, 1% (w/v) [21]. 2.0 ml of fresh starter culturewas inoculated to 50 ml of medium and incubatedat 37°C with continuous agitation at 100 rpm for 6 days. Sampleswere collected at regular intervals, centrifuged at 8000 rpm for 10 min at 4°C and the clear supernatant was used for the assay of enzyme activity.
Chitinase Enzyme Assay
Chitinase activity was determined by measuring the amount of the reducing end groups,liberated in reaction mixture after degradation from acid-swollen and colloidal chitin,in a colorimetric assay using 3,5-dinitrosalicylic acid (DNS) reagent [24]. The chitinase activity was determined by mixing 1 ml crude enzyme solution with 1 ml of acid-swollen chitin (1%, w/v) in 50 mM sodium acetate buffer (pH 5) at 37°C for 1 hour. N-acetylD-glucosamine (1 mg/ml) was used as the standard. Chitinase activity was expressed as µmoles of product formedper ml of enzyme solution per minute (µmoles ml-1 min-1).
Analysis of chitin hydrolysis products
Samples showing maximum activity as crude enzymes were further analyzed through high performance liquid chromatography. 1 ml of crude enzyme was incubated with 1 ml of acid swollen chitin (1%, w/v) prepared in 0.1 M sodium acetate buffer (pH 5) at 37°C for 1 hour. After centrifugation, the reaction mixture was filtered using syringe filter (0.2 µm), and samples were analyzed through dual λ Absorbance detector HPLC (Waters 2487, India). N-acetyl glucosamine (1 mg/ml, 1000 ppm) was taken as standard and samples (20 µl) were analyzed at 210 nm, on C-18 column using acetonitrile: water (70:30) as mobile phase with flow rate of 0.8 ml min-1according to the method described by [25].
RESULTS
A large number of bacterial strains were isolated on nutrient agarplates from samples collected from a variety of seafood wastes and wastewater effluent. The isolates were categorised in five groups based on their source of isolation. Morphological features of the chitinolytic organisms were studied by Grams staining method; microscopic studies confirmed majority of isolates were bacilli or cocci, detailed results for five groups are summarised in (Table 1).
Table 1: Characterization of strains isolated from samples collected from wastewater.
|
Isolate No. |
Nutrient Agar * |
Colloidal-chitin M9 Medium* |
Morphology of isolates |
Gramstain Test (+/-) |
|
PASB 1 |
+ |
- |
White, raised, opaque, rough |
- |
|
PASB 2 |
+ |
+ |
White, raised, opaque, rough |
+ (bacilli) |
|
PASB 3 |
+ |
+ |
White, raised, translucent, smooth |
+ (cocci) |
|
PASB 4 |
+ |
- |
White, raised, opaque, rough |
- |
|
PASB 5 |
+ |
+ |
Orange, raised, opaque, smooth |
+ (cocci) |
|
PASB 6 |
+ |
+ |
Yellow, raised, opaque, smooth |
+ (cocci) |
|
PASB 7 |
+ |
- |
Tan, raised, opaque, rough |
- |
|
PASB 8 |
+ |
- |
White, raised, opaque, smooth |
- |
*+ indicates growth; - indicates no growth
Table 2
Table 2: Characterization of strains isolated from samples collected from Squid (Decapodiformes){9, 10}; and Shrimps(large king prawns) {11-16, 50-55}.
|
Isolate No. |
Nutrient Agar * |
Colloidal-chitin M9 Medium* |
Morphology of isolates |
Gramstain Test (+/-) |
|
PASB 9 |
+ |
+ |
White, raised, translucent, rough |
+ (cocci) |
|
PASB 10 |
+ |
- |
White, raised, translucent, rough |
- |
|
PASB 11 |
+ |
+ |
Yellow, raised, translucent, smooth |
+ (bacilli) |
|
PASB 12 |
+ |
+ |
White, raised, opaque, smooth |
+ (bacilli) |
|
PASB 13 |
+ |
+ |
Yellow, raised, opaque, rough |
+ (bacilli) |
|
PASB 14 |
+ |
+ |
Brown, raised, opaque, rough |
+ (cocci) |
|
PASB 15 |
+ |
- |
Orange, raised, opaque, smooth |
- |
|
PASB 16 |
+ |
- |
White, raised, opaque, smooth |
- |
|
PASB 50 |
+ |
+ |
Orange, raised, opaque, rough |
+ (bacilli) |
|
PASB 51 |
+ |
+ |
Transparent, raised, smooth |
+ (bacilli) |
|
PASB 52 |
+ |
+ |
White, raised, opaque, smooth |
+ (bacilli) |
|
PASB 53 |
+ |
+ |
Transparent, raised, smooth |
+ (bacilli) |
|
PASB 54 |
+ |
+ |
White, raised, opaque, smooth |
+ (bacilli) |
|
PASB 55 |
+ |
+ |
Brown, raised, opaque, smooth |
+ (bacilli) |
*+ indicates growth; - indicates no growth
Table 3
Table 3: Characterization of strains isolated from samples collected from Sea Crab (C. cruciata)
|
Isolate No. |
Nutrient Agar * |
Colloidal-chitin M9 Medium* |
Morphology of isolates |
Gramstain Test (+/-) |
|
PASB 17 |
+ |
- |
White, raised, opaque, smooth |
- |
|
PASB 18 |
+ |
- |
Yellow, raised, opaque, smooth |
- |
|
PASB 19 |
+ |
- |
White, raised, opaque, smooth |
- |
|
PASB 20 |
+ |
+ |
Raised, transparent, smooth |
+ (bacilli) |
|
PASB 29 |
+ |
+ |
Yellow, raised, opaque, smooth |
+ (bacilli) |
|
PASB 30 |
+ |
+ |
Red, raised, translucent, smooth |
+ (bacilli) |
|
PASB 31 |
+ |
- |
White, raised, opaque, rough |
- |
|
PASB 32 |
+ |
- |
White, raised, translucent, smooth |
- |
|
PASB 47 |
+ |
- |
White, raised, opaque, smooth |
- |
|
PASB 48 |
+ |
- |
White, raised, opaque, smooth |
- |
|
PASB 49 |
+ |
- |
Red, raised, opaque, smooth |
- |
|
PASB 56 |
+ |
- |
Yellow, raised, opaque, smooth |
- |
|
PASB 57 |
+ |
+ |
Yellow, raised, opaque, smooth |
+ (bacilli) |
|
PASB 58 |
+ |
+ |
White, raised, opaque, smooth |
+ (bacilli) |
|
PASB 59 |
+ |
- |
Yellow, raised, opaque, smooth |
- |
|
PASB 60 |
+ |
+ |
Transparent, raised, smooth |
+ (bacilli) |
|
PASB 61 |
+ |
+ |
Yellow, raised, opaque, smooth |
+ (bacilli) |
|
PASB 62 |
+ |
+ |
White, raised, opaque, smooth |
+ (bacilli) |
|
PASB 63 |
+ |
- |
Red, raised, opaque, smooth |
- |
|
PASB 64 |
+ |
- |
Transparent, raised, smooth |
- |
|
PASB 65 |
+ |
+ |
Transparent, raised, smooth |
+ (bacilli) |
|
PASB 66 |
+ |
- |
Translucent, raised, rough |
- |
|
PASB 67 |
+ |
- |
Transparent, raised, smooth |
- |
|
PASB 68 |
+ |
+ |
Yellow, raised, opaque, smooth |
+ bacilli |
|
PASB 69 |
+ |
+ |
White, raised, opaque, rough |
+ bacilli |
|
PASB 70 |
+ |
+ |
Yellow, raised, opaque, smooth |
+ bacilli |
|
PASB 71 |
+ |
- |
Transparent, raised, smooth |
- |
*+ indicates growth; - indicates no growth
Table 4
Table 4: Characterization of strains isolated from samples collected from a variety of Fish Scales.
|
Isolate No. |
Nutrient Agar * |
Colloidal-chitin M9 Medium* |
Morphology of isolates |
Gramstain Test (+/-) |
|
PASB 21 |
+ |
- |
White, raised, opaque, rough |
- |
|
PASB 22 |
+ |
- |
Brown, raised, opaque, rough |
- |
|
PASB 23 |
+ |
- |
White, raised, opaque, rough |
- |
|
PASB 24 |
+ |
- |
White, raised, opaque, smooth |
- |
|
PASB 25 |
+ |
+ |
White, raised, opaque, rough |
+ (bacilli) |
|
PASB 26 |
+ |
- |
Cream, raised, opaque, smooth |
- |
|
PASB 27 |
+ |
- |
Transparent, raised, rough |
- |
|
PASB 28 |
+ |
- |
Yellow, raised, translucent, smooth |
- |
*+ indicates growth; - indicates no growth
Table 5
Table 5: Characterization of strains isolated from samples collected from Prawn shells.
|
Isolate No. |
Nutrient Agar * |
Colloidal-chitin M9 Medium* |
Morphology of isolates |
Gramstain Test (+/-) |
|
PASB 33 |
+ |
+ |
Red, raised, opaque, smooth |
+ (bacilli) |
|
PASB 34 |
+ |
- |
White, raised, opaque, smooth |
- |
|
PASB 35 |
+ |
- |
Orange, raised, opaque, smooth |
- |
|
PASB 36 |
+ |
+ |
Cream, raised, opaque, smooth |
+ (bacilli) |
|
PASB 37 |
+ |
- |
Transparent, raised, smooth |
- |
|
PASB 38 |
+ |
+ |
Orange, raised, opaque, smooth |
+ (cocci) |
|
PASB 39 |
+ |
- |
Yellow, raised, opaque, smooth |
- |
|
PASB 40 |
+ |
- |
Red, raised, opaque, smooth |
- |
|
PASB 41 |
+ |
- |
White, raised, opaque, rough |
- |
|
PASB 42 |
+ |
- |
White, raised, opaque, smooth |
- |
|
PASB 43 |
+ |
- |
Cream, raised, opaque, rough |
- |
|
PASB 44 |
+ |
+ |
Yellow, raised, opaque, smooth |
+ (bacilli) |
|
PASB 45 |
+ |
- |
Transparent, raised, smooth |
- |
|
PASB 46 |
+ |
- |
White, raised, opaque, smooth |
- |
*+ indicates growth; - indicates no growth
These isolates were subjected to a process for screening of chitinolytic strains by spot-inoculating on plates of a minimal medium containing colloidal chitin (CCM). In this repeated screening method using 71 different bacterial isolates, only 32 were found to be chitinase positive. Pictures of CCM plates showing chitinase positive bacterial colonies are presented in (Figure 1 A-C).
Figure 1: Pictures of CCM plates showing chitinase positive bacterial colonies.
All 32 chitinase positive isolates were further taken for a secondary screening in production medium for enzyme production and samples at regular interval were assyed for chitinase units produced by isolates, the results for are presented in (Figure 2)
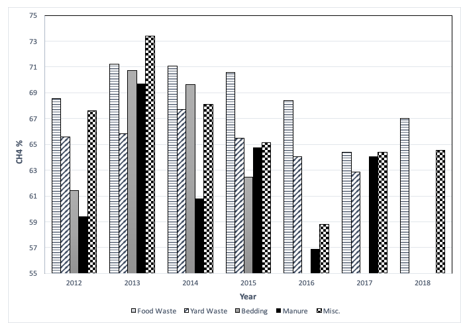
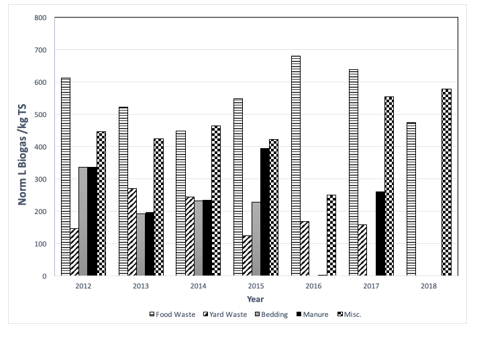
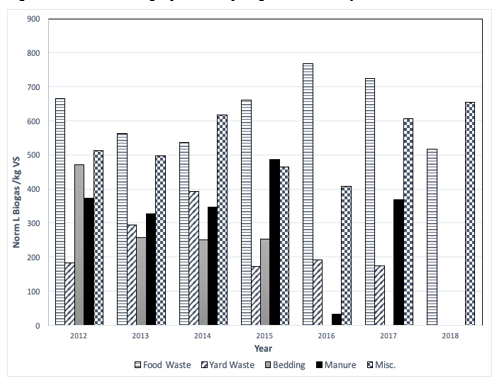
Figure 2: Enzyme production and samples at regular interval.
and (Table-6).
Table 6: ChitinaseEnzyme synthesis by isolated bacterial cultures.
|
Isolate source/name |
Extracellular Chitinase produced (micromoles/ ml.min) |
|
Wastewater/PASB 2 |
2.5 |
|
Wastewater/PASB 3 |
1.43 |
|
Wastewater/PASB 5 |
2.15 |
|
Wastewater/PASB 6 |
5.01 |
|
Shrimp PASB 9 |
8.98 |
|
Shrimp PASB11 |
25.61 |
|
Shrimp PASB12 |
31.64 |
|
Shrimp PASB13 |
3.65 |
|
Shrimp PASB14 |
35.41 |
|
Shrimp PASB50 |
7.46 |
|
Shrimp PASB51 |
30.5 |
|
Shrimp PASB52 |
5.86 |
|
Shrimp PASB53 |
8.88 |
|
Shrimp PASB54 |
19.979 |
|
Shrimp PASB55 |
12.5 |
|
Prawn PASB36 |
1.56 |
|
Prawn PASB38 |
26.34 |
|
Prawn PASB44 |
7.56 |
|
Crab PASB29 |
4.33 |
|
Crab PASB30 |
2.24 |
|
Crab PASB57 |
25.588 |
|
Crab PASB58 |
8.65 |
|
Crab PASB60 |
5.65 |
|
Crab PASB61 |
2.12 |
|
Crab PASB62 |
8.23 |
|
Crab PASB65 |
4.23 |
|
Crab PASB68 |
8.34 |
|
Crab PASB69 |
4.37 |
|
Crab PASB70 |
5.33 |
|
Fish PASB25 |
6.85 |
Thehigher enzyme producers strains (PASB-11, 12, 14, 51, 54 and 55) were isolated from samples of Shrimps (large king prawns), PASB-38 was obtained from prawns;their morphology results are presented in tables 2 and 4.The 8thgood chitinase producer designated as PASB-57 was isolated from sea crab shells (Charybdis cruciate), morphology results are presented in table-3.
Higher Chitinase enzyme producers PASB-11, 12, 14, 38, 51, 54, 55, 57 with enzyme units 25.61, 31.64, 35.41, 26.34, 30.5, 19.9, 12.5 and 25.6 IU, respectively were studiedfor theirHPLC profile (Figure 3).
Figure 3: Higher Chitinase enzyme producers PASB-11, 12, 14, 38, 51, 54, 55, 57 with enzyme units.
These higher chitinase producers were shortlisted for identification through 16sRDNA sequencing followed by analysis through BLASTn (NCBI), CLUSTAL W (EBI) and SEQMATCH (RDP database). The results suggested the strains as Acinetobactor spp. (PASB 14), Bacillus spp. (PASB-51), Kurthiagibsonii (PASB38; 54) and Alcaligenes spp. (PASB 11; 12; 55 and 57).
DISCUSSIONS
Microbial production of chitinase has captured the worldwide attention of both industrial and scientific environments, not only because of its wide spectrum of applications but also for the lacuna of an effective production method. A wide range of microbes capable of producing chitinases have been isolated from different sources such as soil, peanut hulls, marine waste, etc.In the present study chitinase producing microorganisms were isolated from seafood wastes shrimp shells, prawn shells, crab shells and other marine samples. Around 32 chitin degrading organisms were isolated, out of which four potential chitinolytic bacteria were selected for further studies based on their high extracellular chitinase production. Their identification and characterization was done through 16S rDNA sequencing followed by analysis using various bioinformatics tools. The four isolates were identified as Bacillus spp., Kurthiagibsonii andAlcaligenes spp. Earlier studies have reported the chitinase production by Alcaligenes faecalis AU02 to be maximum (258 U/ mL) after 48 h at 37°C in a medium containing 1% shrimp and crab shell powder in basal medium (pH 8.0) [26]. Bacillus sp. TKU006 has been reported to produce protease and chitinase on shrimp shell powder and crab shell powder at 25°C for 2 days [27]. The chitinolytic activity in the culture supernatant of Bacillus pumilus reached a maximum of 79.8 U/100 ml after 8 days of incubation using basal medium [28]. In comparison to these findings, our results show an increased enzyme activity of 30.502 mmoles min-1ml-1,19.979 mmoles min-1 ml-1, 12.5 mmoles min-1ml-1 and 25.588 mmoles min-1ml-1 for Bacillus spp. PASB-51, Kurthiagibsonii PASB-54, Alcaligenes spp. PASB-55 and PASB-57, respectively [29].
First reported that the chitinolytic activity of Kurthia gibsonii Mb126 strain isolated from prawn shell powder could be increased up to 65 U/ml under optimum conditions. They further purified the enzyme through four steps including ammonium sulphate precipitation, affinity adsorption, ion exchange chromatography and gel filtration chromatography. The chitinase was purified 16.11-fold through Sephadex G 100 gel filtration. The specific activities of the crude and purified enzyme were 0.64 U/mg and 10.31 U/mg proteins, respectively. The enzyme was most active at pH 6.5 at 40°C [30]. In our study, the chitinolytic activity of Kurthiagibsonii (PASB-54) has been found to be 19.979 mmoles/ min/ml. A qualitative HPLC analysis successfully detected the presence of monomer NAG, a part of chitin hydrolysis product, thereby proving chitinase production in the medium by these four strains. Also, these isolates were capable of degrading chitin to its monomer unit NAG.
With respect to present results and comparison with the best knowledge about other chitinase producers, these isolates may have the capability for production of chitinase on an industrial scale. Also, many thermostable chitinases have been identified from a variety of microorganisms. The chitinolytic microbes identified in this study can be further analyzed for their stability over high temperatures and thus could prove useful for other biotechnological applications.
ACKNOWLEDGMENTS
We are thankful to National Centre for Cell Science (NCCS), Pune, India for providing the sequencing facility. We also acknowledge Meenal Rastogi for valuable discussions and feedback.
REFERENCES
- Tang, WJ, Fernandez, JG; Sohn, JJ; Amemiya, CT (2015). Chitin is endogenously produced in vertebrates. Curr Biol. 25: 897-900.
- Flach J, Pilet PE, Jolles P. What's new in chitinase research? Experientia. 1992; 48: 701-716.
- Patil RS, Ghormade V, Deshpande MV. Chitinolytic enzymes: an exploration. Enzyme Microb Technol. 2000; 26: 473-483.
- Kuddus SM, Ahmad R.I.Z. Isolation of novel chitinolytic bacteria and production optimization of extracellular chitinase. Journal of Genetic Engineering and Biotechnology. 2013; 1: 39-46.
- Wang SL and Chio SH. Deproteinization of shrimp and crab shell with the protease of Pseudomonas aeruginosa K-187. Enzyme Microb. Technol 1998; 629-633.
- Stoykov YM, Pavlov AI, Krastanov AI. Chitinase biotechnology: Production, purification, and application. Engg in Life Sci. 2015. 15; 30-38.
- Carroad PA and Tom RA. Bioconversion of shellfish chitin wastes: process, concept and selection of microorganisms. J Food Sci. 1978; 43: 1158-1161.
- Suresh PV and Chandrasekaran M. Utilization of prawn waste for chitinase production by the marine fungus Beauveriabassiana by solid state fermentation. World J Microbiol Biotechnol. 1998; 14: 655-660.
- Ifuku, S (2014). "Chitin and Chitosan Nanofibers: Preparation and Chemical Modifications". Molecules. 19 (11): 18367-18380.
- Mathivanan N, Kabilan V, Murugesan K. Purification, characterization and antifungalactivityof chitinase form Fusariumchlamydospourm, a mycoparasite to groundnut rust, Pucciniaarachidis. Can J Microbiol 1998; 44: 656-651.
- Cody RM. Distribution of chitinase and chitobiase in Bacillus. CurrMicrobiol 1989; 19: 201-205.
- Bhattacharya D, Nagpure A, Gupta RK. Bacterial chitinases: properties and potential. Critical Reviews in Biotechnology. 2007; 27: 21-28.
- Swiontek Brzezinska M, Jankiewicz U, Burkowska A, Walczak M. Chitinolytic Microorganisms and Their Possible Application in Environmental Protection. Current Microbiology. 2014; 68: 71-81.
- Gohel V, Maisuria V, Chhatpar HS. Utilization of various chitinous sources for production of mycolytic enzymes by Pantoeadispersa in bench top fermenter. EnzymeMicrobialTech. 2007; 40: 1608-1614.
- Pichyangkura R, Kudan S, Kultiyawong K, Sukwattanasinitt M, Aiba S. Quantitative production of 2-acetamido-2-deoxy-D-glucose from crystalline chitin by bacterial chitinase. Carbohydr Res. 2002; 337: 557-559.
- Vyas P and Deshpande MV. Enzymatic hydrolysis of chitin by Myrothecium verrucariachitinase complex and its utilization the produce SCP. J Gen ApplMicrobiol. 1991; 37: 267-275.
- Cody RM, Davis ND, Lin J, Shaw D. Screening microorganisms for chitin hydrolysis and production of ethanol from amino sugars. Biomass. 1990; 21: 285-295.
- Abd-Aziz S, Fernandez CC, Md. Salleh MM, Illias RM, Hassan MA. Effect of agitation and aeration rates on chitinase production using Trichodermavirens UKM1 in 2-l stirred tank reactor. Appl Biochem Biotechnol. 2008; 150(2): 193-204.
- Shah JM, Raghupathy V, Veluthambi K. Enhanced sheath blight resistance in transgenic rice expressing an endochitinase gene from Trichodermavirens. Biotechnol Lett. 2009; 31: 239-244.
- Nagpure A, Choudhary B, Gupta RK. Chitinases: in agriculture and human healthcare. Crit Rev Biotechnol. 2014; 34: 215-232.
- Ghosh U, Chakrobarty S. Segregation and categorization of chitinase producing bacteria using exoskeleton of Penaeusindicusas a substrate from Sagar Island, Sunderbans. International Journal of Chemical and analytical science 2010; 1: 193-194.
- Bartholomew JW and Mittwer T. THE GRAM STAIN. Bacteriol Rev. 1952; 16: 1-29.
- Annamalai N, Giji S, Arumugam M, Balsubramanian T. Purification and characterization of chitinase from Microccous sp.AG84 isolated from marine environment. African Journal of Biotechnology Research. 2010; 4: 2823.
- Miller GL. Use of Dinitrosalicylic acid reagent for determination of reducing sugars. Ana Chem. 1959; 31: 426-428.
- El-Dein A, Hosny MS, El-Shayeb NA, Abood A, Abdel-Fattah AM. A Potent Chitinolytic Activity of Marine Actinomycete sp. and Enzymatic Production of Chitooligosaccharides. Australian Journal of Basic and Applied Sciences. 2010; 4: 615-623.
- Annamalai N, Veeramuthu Rajeswari M, Vijayalakshmi S, Balasubramanian T. Purification and characterization of chitinase from Alcaligenesfaecalis AU02 by utilizing marine wastes and its antioxidant activity. Annals of Microbiology. 2011; 61: 801-807.
- Wang SL, Chao CH, Liang TW, Chen CC. Purification and characterization of protease and chitinase from Bacillus cereus TKU006 and conversion of marine wastes by these enzymes. Mar Biotechnol (NY) 2009; 11: 334-344.
- Tasharrofi N, Adrangi S, Fazeli M, Rastegar H, Khoshayand MR, Faramarzi MA. Optimization of Chitinase Production by Bacillus pumilus Using Plackett-Burman Design and Response Surface Methodology. Iran J Pharm Res. 2011; 10: 759-768.
- Paul MK, Mini KD, Antony A, Radhakrishnan EK, Mathew J. Utilization of prawn shellpowder for the production of chitinase by Kurthia gibsonii MB126. International Journal of Pharma and Bio Sciences. 2012; 3: 163-172.
- Paul MK, Mini KD, Mathew J. Purification and characterization of chitinase enzyme from Kurthiagibsonii Mb126. Journal of Environment and Biotechnology Research. 2012; 2: 37-44.