Mutant of RNA Binding Protein Zc3h14 Causes Cell Growth Delay or Arrest through Inducing Multinucleation and DNA Damage
- 1. The National Institute of Biological Sciences, Department of Neurology, China
- 2. The Department of Neurology, Beth Israel Deaconess Medical Center and Harvard Medical School, USA
Abstract
ZC3H14 proteins bind poly (A) tail of RNAs through the CCCH (5) zinc finger domain, shuttle between the nucleus and the cytoplasm, and play roles in nuclear exporting and proper polyadenylation of mRNA transcripts. However, molecular and cellular mechanisms of mammalian ZC3H14 remain largely unknown. Here we identified the mouse Zc3h14 gene and its encoded Zc3h14 protein. We found that Mouse Zc3h14 gene had at least two transcript isoforms, both were widely expressed in various tissues. Mouse ZC3H14 protein had one N-terminus Q-rich domain and one CCCH (5) zinc fingers domain. Both the two isoforms localized to the nucleus, and intriguingly, formed multiple nuclear speckles. We mapped the core nuclear localization signal to 292-RKRK sequences. Serial mutation analysis showed that the CCCH (5) zinc fingers domain was necessary but not sufficient for nuclear speckle formation. Over-expression of CCCH(5) domain as a dominant negative mutant, caused dramatic cell cycle delay, accompanied with multiple morphological abnormalities, such as long processes growing-out, multinucleation, and nuclear fragmentation. Further studies demonstrated that the CCCH (5) domain induced nuclear foci formation of the DNA damage marker g-H2AX. RT-PCR analysis showed that a group of DNA damage/repair genes, such as Cdk2, Ddb1, Mdm2, p21, p53, Rad50, as well as two RNA binding proteins, Pabpc1 and Rbm10, were dramatically up-regulated. Immunoprecipitation and mass spectrometry analysis further showed that the CCCH (5) domain could bind with RNA binding protein RBM10 in an RNA-dependent manner, and the CCCH (5) domain could also immunoprecipitate a group of chromatin stability/DNA damage-related proteins. In brief, our data for the first time demonstrated that the RNA binding protein, ZC3H14, is associated with genome/chromatin stability and DNA damage/repair.
Keywords
ZC3H14; RNA binding proteins; Cell growth delay; DNA damage and repair; gamma-H2AX; p53; RBM10.
CITATION
Hu J, Gao S (2014) Mutant of RNA Binding Protein Zc3h14 Causes Cell Growth Delay/Arrest through Inducing Multinucleation and DNA Damage. JSM Biochem Mol Biol 2(1): 1006.
ABBREVIATIONS
AAs, amino acids; DAPI, 4’-6-Diamidino-2-phenylindole; DSBs, DNA double-strand breaks; EGFP, enhanced green fluorescent protein; FBS, fetal bovine serum; γ-H2AX, gamma H2AX; mRNP, mRNA ribonucleoprotein; MW, molecular weight; NLS, nuclear localization signal; ORF, open reading frame; PFA, paraformaldehyde ; Pabs, Poly(A) binding proteins; PHDL cells, pancreatic PNA-HSA double-low cells; PI, isoelectric point; RRM, RNA recognition motif; RT-PCR, retrotranscribed polymerase chain reaction; WT, wild type
INTRODUCTION
Message RNA (mRNA) transcripts are coated from cap to tail with RNA binding proteins, which are involved in dynamically processing, packaging, exporting and translational modulation of the transcripts [1-4]. These proteins also regulate the fate of the mature mRNA transcripts. RNA binding proteins in mRNP complexes act as posttranscriptional activators and repressors of mRNA translation and degradation [4-6]. Poly (A) binding proteins (Pabs) are one sub-family of RNA binding proteins. They play key roles in regulating posttranscriptional mRNA translation and degradation. Two types of poly(A) binding motifs have been identified: the RNA recognition motif (RRM) and the CCCH zinc fingers motif [7,8]. Different RRMs in one protein can each bind RNA with varying specificity and affinity [7-9]. Different CCCH domains relate to the zinc-binding motifs of the largest subunit of RNA polymerases I, II, and III [10]. CCCH zinc fingers can bind to poly(A), poly(G), poly(U), single-stranded DNA [10], and may bind to RNA alone or coordinately with Arginine–glycine–glycine (RGG) repeat motif [10-12]. Different CCCH-type proteins can interact with the 3’-untranslated region of various mRNAs [13]. ZC3H14 protein is a subclass of ploy(A) binding proteins, which contains five or seven tandem CCCH zinc finger (CCCH) repeats but lacks RRM motif [10-12]. Nab2 is the orthologue of human and mouse ZC3H14 proteins. ZC3H14/Nab2 shuttles between the nucleus and the cytoplasm and is required for both nuclear export and proper polyadenylation of mRNA transcripts [10,14]. They contain five (human and mouse) or seven (yeast) CCCH zinc finger repeats at their C-terminus (CCCH(5) or CCCH(7)). ZC3H14 binds poly(A) specifically through the CCCH zinc fingers [12]. Most recently, it was reported that ZC3H14 played roles in neural system and immune system [15-18]. There was a study reported the structural basis for polyadenosine-RNA binding by ZC3H14/Nab2 Zn fingers and its function in mRNA nuclear export [19]. However, the molecular and cellular mechanisms for physiology and pathological roles of mammalian ZC3H14 protein remain largely unknown.
In this study, we characterized mouse Zc3h14 gene and encoded Zc3h14 protein. We first detected the expression pattern and transcript isoforms of Zc3h14 gene. Then we determined the sub-cellular localization using the EGFP or mCherry tagged protein and found that mouse ZC3H14 protein localized to the nucleus and formed nuclear speckles. We further mapped the core nuclear localization signal (NLS) of Zc3h14 to 292-RKRK sequences. Functional studies showed that the ZC3H14 CCCH(5) domain expression caused dramatic cell growth delay, accompanied with long process growing-out, multinucleation, and nuclear gfragmentation. Unexpectedly, we found that overexpression of the ZC3H14 CCCH(5) domain caused DNA damage (marker γ-H2AX foci formation) and would induce dramatic up-regulation of a group of DNA damage/repair genes. We further performed biochemistry analysis and found that Zc3h14 binds a group of RNA binding proteins, which were associated with genome stability and DNA damage/repair. Our work for the first time showed that the RNA binding protein Zc3h14 is also related with genome/chromatin stability.
MATERIALS AND METHODS
Tissue isolation and Cell culture
Adult ovaries were isolated from C57BL/6J mice. All animal studies adhered to procedures consistent with the National Institute of Biological Sciences Guide for the care and use of laboratory animals. Pancreatic PNA-HSA double-low (PHDL) epithelial cells were derived from fetal pancreas of mice (unpublished data). PHDL cells were cultured under the conditions modified from the previous report [20]. In brief, the pancreatic PHDL cells were cultured in DMEM based medium, which contained 2.5% FBS (Hyclone), 2mM glutamine, 100IU/ml penicillin and 100ug/ml streptomycin. P19 cells and 293T cells were cultured in DMEM based medium, which contained 10% FBS (Hyclone), 2mM glutamine, 100IU/ml penicillin and 100ug/ ml streptomycin. Single cell colony formation was analyzed by limited-dilution. Cell number was counted at different time points (see below).
cDNA cloning, Vector construction, lentivirus production and infection
The mouse Zc3h14 ORF was obtained by RT-PCR from adult ovary and PHDL cells (See Table 1 to find the primers used).
Table1: The primers had been used for cloning full-length ORFs and mutants.
|
Primers Fragments |
forward: 5' to 3' |
reverse: 5' to 3' |
Products size (bp) |
|
Zc3h14-ORF, Isoform 1 |
atggaaatcggcaccgagatcag |
ctcactgctctgaggtcgaatccat |
2205 |
|
Zc3h14-ORF, Isoform 2 |
atggaaatcggcaccgagatcag |
ctcactgctctgaggtcgaatccat |
1815 |
|
Zc3h14-N590 |
atggaaatcggcaccgagatcag |
cacactcaggtcagtcatctcggc |
1570 |
|
Zc3h14-N303 |
atggaaatcggcaccgagatcag |
aacaaccgaacttaccacaggcaat |
909 |
|
Zc3h14-CCCH(5) |
gagatgactgacctgagtgtggcac |
ctcactgctctgaggtcgaatccat |
456 |
|
Zc3h14-292-RKRK to 292-PNTR mutation, N-terminus |
atggaaatcggcaccgagatcag |
cactacgcgtgtttggaaagttttcgtcctcaatactc |
885 |
|
Zc3h14-292-RKRK to 292-PNTR mutation, C-terminus |
cactacgcgtttgcctgtggtaagttcggttgtta |
ctcactgctctgaggtcgaatccat |
1326 |
The ORF and mutants were cloned into pEGFP-N1 vector for transient expression. They were cloned into pWPI lentivirus vector for stable expression. The lentivirus was produced by co-transfection of the lentivirus vector and two helper vectors, pSPAX2 and pMD2.G [21]. Vigofect reagent was used according to the protocol provided by the manufacturer (Vigorous).
RT-PC
Total RNA samples were prepared from adult brain, heart, kidney, liver, lung, pancreas, skin, spleen, ovary, testis, PHDL cells and P19 cells. The RNA was reverse-transcribed by M-MLV reverse transcriptase (Promega) and amplified by PCR for 30 cycles. PCR cycling was performed at 98°C for 2 min followed by 98°C for 15sec, 54°C to 60°C for 15sec, 72°C for 1min, and finally 72°C for 8min, by using the PrimeSTAR HS DNA polymerase (Takara). (See Table 2 to find the primers used).
Table 2: The primers had been used for RT-PCR analysis.
|
Primers genes |
Forward: 5' to 3' |
Reverse: 5' to 3' |
Products size (bp) |
|
Zc3h14 |
acggttcttctcgcccttctat |
ttgcctctgttgagttccttga |
683 |
|
Cdk2 |
ccggctcgacactgagactgaa |
gtgaaggacacggtgagaatgg |
271 |
|
Chek1 |
catcaggtggtatgtcagagtc |
cccttagaaagtcggaagtcaa |
502 |
|
Ddb1 |
tggtgttgtatttgtcgggtct |
gatgcctggtaagtcaatgctg |
265 |
|
Mdm2 |
gcaagcacctcacagattccag |
gatccttcagatcactcccacc |
364 |
|
p21 |
gtgagcagttgcgccgtgattg |
tgagggctaaggccgaagatgg |
478 |
|
p53 |
gggacagccaagtctgttatg |
ggagtcttccagtgtgatgat |
432 |
|
Pabpc1 |
actacaagcccaccaagcgaaag |
gatgtcccaagctattccacagtaa |
420 |
|
Rad50 |
gtttactcccagttcattacgc |
ttcctgctcttctatgtcattcttt |
324 |
|
Rbm10 |
cggtgggcacgcagtctcaata |
ttcctccgtttgggctctggtg |
542 |
|
b-Actin |
accaactgggacgacatggagaaga |
ctctttgatgtcacgcacgatttc |
414 |
Immunofluorescent staining and confocal microscop
Cells growing on cover-slips were fixed with 4% PFA for 20min. The samples were permeabilized with 0.5% triton X-100 and blocked with 5% normal horse serum for 2h. The anti-γ H2AX antibody was incubated overnight at 4°C. The Alexa-594 conjugated secondary antibodies were incubated for 1h at room temperature. Samples were further counterstained with 100ng/ ml DAPI. Images were obtained with Zeiss LSM510 Meta confocal microscope.
Immunoprecipitation and western blottin
293T cells were transfected to transiently express Zc3h14 FLAG-EGFP, Zc3h14(N590)-FLAG-EGFP, Zc3h14(CCCH(5)) FLAG-EGFP or FLAG-EGFP. Samples were collected 48 hours after transfection. Immunoprecipications were performed according to the protocol that provided by the manufacturer (FLAGIPT-1; Sigma). To digest RNA, RNase A was added to the samples at a final concentration of 50 µg/ml. The mixture was incubated for 20 min at 37°C before the final elution. Proteins were separated in 10% SDS-polyacrylamide gels and transblotted to PVDF membrane. Immunoblot analysis was performed with anti-FLAG monoclonal antibody (Sigma) and anti-RBM10 polyclonal antibody (Aviva). Blots were detected by using ECL (Amersham Biosciences) according to the protocol that provided by the manufacturer.
Silver staining and mass spectrometry analysi
Proteins were separated in 10% SDS-polyacrylamide gels. Silver staining was performed according to the protocol that provided by the manufacturer (PROTSIL2-1KT, Sigma). The bands of interest were analyzed by LTQ linear ion trap mass spectrometer (Thermo Electron).
RESULTS
The expression pattern, sequence analysis and construct design of mouse Zc3h14 gene
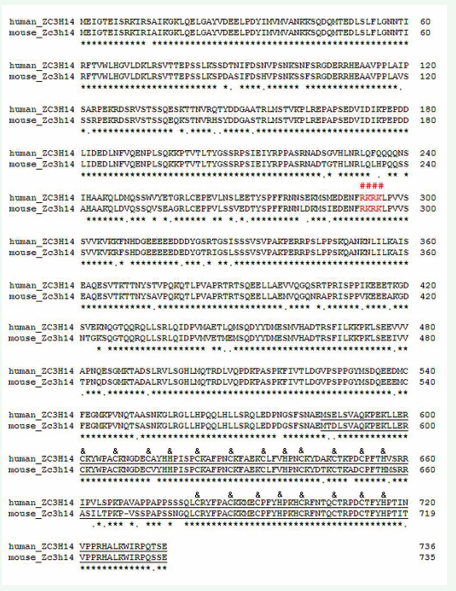
The human and mouse ZC3H14 had 89.7% identities in amino acid sequences in their long form (Figure 1).
Figure 1: Sequence alignment between the human and mouse ZC3H14 proteins: The long form of human and mouse ZC3H14 have 89.7% identities in amino acid sequences. The predicted nuclear localization signal sequences, 292-RKRK, are well conserved (red colored and with “#” above them. The sequences of CCCH(5) domain are underlined and the CCCH core amino acids are labeled with “&” above them.
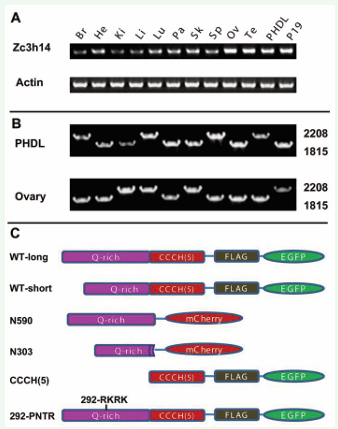
The protein contained one Q-rich domain in N-terminus and one CCCH zinc fingers domain in C-terminus (Figure 1). RT-PCR results showed that the Zc3h14 gene was expressed in all the tissues analyzed. It was highly expressed in ovary and testis (Figure 2A). Two transcript isoforms had been cloned from pancreatic PHDL cells and ovary by using the primers list in table 1. The long form ORF was 2208bp in length. It encoded a 735 AAs long polypeptide (predicted MW: 82.4 kD, PI: 7.39). The short form ORF was 1815bp in length. It encoded a 604 AAs long polypeptide (predicted MW: 67.9 kD, PI: 8.14). The two transcript isoforms had similar expression levels in PHDL cells and ovary (Figure 2B). To characterize the functions of the different domains and to map the core nuclear localization signal sequences, partial deletion mutations and point mutations were designed (Figure 2C).
Figure 2: Expression pattern, and constructs design of mouse Zc3h14 gene. (A) RT-PCR analysis. Zc3h14 is expressed in all the tissues and cell lines that having been detected (Br, Brain; He, Heart; Kidey, Ki; Li, Liver; Lu, Lung; Pa, Pancreas; Sk, Skin; Sp, Spleen; Ov, Ovary; Te, Testis; PHDL, pancreatic PNA-HSA double-low epithelial cells; P19, P19 embryonal carcinoma cells). Zc3h14 is highly expressed in ovary and testis. (B) Two transcript isoforms are found in ovary and PHDL cells. The two transcript isoforms have similar expression levels in PHDL cells and ovary. (C) Constructs have been designed for localization, biochemistry and functional studies. (WT: wild type; N590 and N303: N-terminus that including the first 590 or 303 AAs of the long form, respectively; CCCH(5): the five CCCH zinc fingers; 292-PNTR: the 292-RKRK sequences are mutated to 292-PNTR).
ZC3H14 localized to the nucleus and formed nuclear speckles
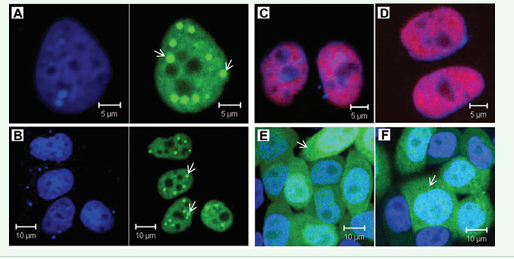
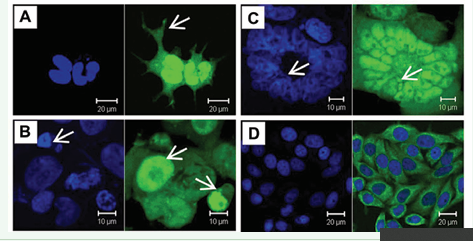
Wild type and mutated ZC3H14 were fuse-expressed with FLAG-EGFP or mCherry. Transient expression showed that ZC3H14 localized to the nucleus (Fiure. 3A, B). Intriguingly, ZC3H14 formed multiple nuclear speckles (Figure 3A, B, arrows). These data suggested that the ZC3H14 protein might play some special roles in DNA and/or RNA regulation. To map the nuclear localization signal of ZC3H14, mutants were fuse-expressed with FLAG-EGFP or mCherry. The results showed that, both the N590 and the N303 mutants localized to the nucleus. However, they did not form nuclear speckles (Figure 3C, D). The CCCH(5) domain localized to both the nucleus and the cytoplasm, but also lost the nuclear speckle formation capacity (Figure 3E). Further analysis suggested that the 292-RKRK sequences maybe the core NLS of mouse ZC3H14 (Figure 1) [22]. Thus we designed a 292-RKRK to 292-PNTR mutant to characterize the NLS (Figure 2C). The result showed that the 292-PNTR ZC3H14 lost its nuclear specific targeting (Figure 3F).
Figure 3: Nuclear localization and NLS mapping of mouse ZC3H14 protein. (A) and (B) Both the long form (A) and short form (B) of wild type ZC3H14 localize to the nucleus. Intriguingly, both of them form multiple nuclear foci (arrows). (C) and (D) Both the N-terminus 590 amino acids and 303 amino acids mutants (N590, C and N303, D) localize to the nucleus. However, these mutants lose the nuclear foci formation capacity. (E) The CCCH(5) domain localize to both the nucleus and the cytoplasm and do not form nuclear foci. (F) The predicted core NLS sequences 292-RKRK are mutated to 292-PNTR. The 292-PNTR ZC3H14 loses its nuclear specific targeting. (green: EGFP; red: mCherry; blue: DAPI).
The CCCH(5) mutant of ZC3H14 caused dramatic cell growth delay
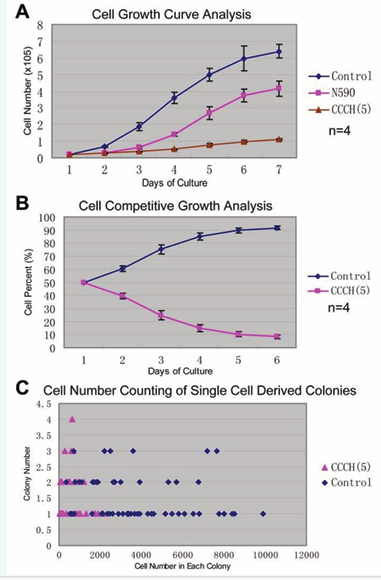
A series of mutants were designed to characterize the functions of mouse ZC3H14. It was reported that the CCCH(5) zinc fingers domain of human ZC3H14 bound poly(A). The N590 and the CCCH(5) mutants were designed as dominant negative mutants. These mutants were fuse-expressed with FLAG-EGFP in pWPI lentivirus vectors (Figure 2C). Stably expressing cells were used for cell growth analysis. We found that both the N590 and the CCCH(5) mutants of ZC3H14 caused cell growth delay in PHDL cells (Figure 4). Cell growth curve analysis showed that the CCCH(5) mutant caused severer cell growth delay than the N590 mutant (Figure 4A). Competitive growth analysis (co-culture of mCherry control cells and CCCH(5)-FLAG-EGFP cells) showed that the cell growth rate was substantially slowed down in the CCCH(5) expressing cells (Figure 4B). Cell number counting of single-cell-derived colonies showed that most of the cells in the CCCH(5) group formed smaller colonies than the cells in the control group (Figure 4C). These results demonstrated that the cell growth rate was severely delayed by the CCCH(5) domain in PHDL cells. Similar results were obtained in P19 embryonal carcinoma cells (data not shown here).
Figure 4: Mutant forms of mouse ZC3H14 caused cell cycle delay. (A) Cell growth curve analysis. Both the N590 and CCCH(5) mutants of ZC3H14 caused cell cycle delay. The CCCH(5) domain induced severer cell cycle delay than the N590 mutant. (B) Competitive growth analysis. Equal number of cells were seeded from mCherry control groups and CCCH(5)-FLAG-EGFP groups. Cells were analyzed by flow-cytometry. The proportion of CCCH(5) stably expressing cells decreased rapidly. (C) Cell number counting of single cell derived colonies after 8 days culture. Most of the cells in the CCCH(5) group formed smaller colonies than those in the control group.
The CCCH(5) mutant of ZC3H14 induced morphological abnormalities and DNA damage
Morphological changes were obviously observed in CCCH(5) stably expressing PHDL cells. Most of the cells lost the irregular polygonal morphology. Many cells grew out long processes, formed multiple nuclei and nuclear fragmentation, and turned round in shape found (Figure 5).
Figure 5:Morphological abnormalities were induced by over-expression of the CCCH(5) domain in PHDL cells. (A) Most cells lost the polygonal epithelial characteristics and grew out long processes (arrow). (B) Multiple nuclei were easily found. One typical cell contained tens of nuclei. Some of the nuclei were fragmented (arrow). (C) Round-shaped cells were easily found (arrows). (D) PHDL cells in the control group showed typical polygonal morphology. (green: EGFP; blue: DAPI).
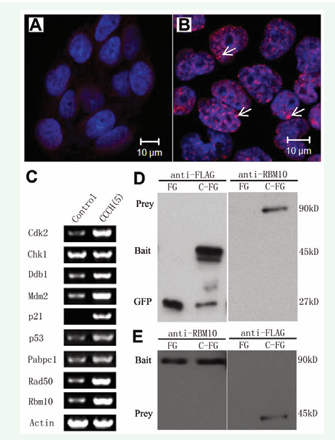
The morphological changes highly suggested that the nuclear/chromatin abnormalities. Since DNA damage was one of the common reasons for cell cycle delay and cell apoptosis [23,24] we further analyzed whether the phenotypes were caused by DNA damage. We stained the CCCH(5) stably expressing cells with DSBs marker γ-H2AX [25]. The results showed that most of the cells were stained highly by γ-H2AX, with strong multiple foci formation in the nucleus (Figure 6A, B). These data demonstrated that DNA damage had been induced by the CCCH(5) domain. To confirm the DNA damage, a group of DNA damage-related genes were analyzed by RT-PCR. The results showed that the expression levels of Cdk2, Ddb1, Mdm2, p21, p53 and Rad50 genes were dramatically up regulated (Figure 6C). The RT-PCR results indicated that DNA damage had been induced by the CCCH(5) mutant.
Figure 6: Over-expression of ZC3H14 CCCH(5) domain induced DNA damage. (A) and (B) Immunofluorecent staining of the DNA double strand breaks (DSBs) marker γ-H2AX. The γ-H2AX signal was very weak in the control group (A). Multiple γ-H2AX nuclear foci were found in most of the CCCH(5)-FLAG-EGFP over-expressed PHDL cells (B) (arrows). (C) RT-PCR analysis. Multiple DNA damage/repair related genes were analyzed. Compared to the control group, expression levels of Cdk2, Ddb1, Mdm2, p21, p53, and Rad50 were dramatically up-regulated. Two apoptosis related genes, Pabpc1 and Rbm10, were also dramatically up-regulated. (D) Western blotting analysis of anti-FLAG immunoprecipitates with anti-RBM10 antibody. (left panel: anti-FLAG; right panel: anti-RBM10; FG and C-FG: lysates prepared from FLAG-EGFP and CCCH(5)-FLAG-EGFP transiently expressed 293T cells, respectively). (E) Western blotting analysis of anti-RBM10 immunoprecipitates with anti-FLAG antibody. (left panel: anti-RBM10; right panel: anti-FLAG; FG: FLAG-EGFP; C-FG: CCCH(5)-FLAG-EGFP). Both CCCH(5)-FLAG-EGFP and RBM10 were detected in corresponding immunopreciptates that untreated with RNase A. (green: EGFP; red: Alexa-594; blue: DAPI).
The CCCH(5) mutant of ZC3H14 interacted with RBM10 and a group of RNA binding proteins, as well as DNA damage/repair related proteins
To find out the potential interactors of Zc3h14 protein that may contribute to its DNA damage effects, immunoprecipitation was performed by using the Zc3h14 CCCH(5) domain as the bait protein and by using the 293T cell lysate a the prey source. Mass spectrometry analysis of the immunoprecipitates was performed and the results demonstrated binding of CCCH(5)-FLAG-EGFP to a group of genome stability associated RNA binding proteins, such as DHX9, HNRNPF, HNRNPH1, HNRNPL, HNRNPR, PABPC1, PABPC4, PABPN1, RBM10, AIFM2, DDB1, FXR1, RAD50, and YWHAZ (See Table 3).
Table 3: Some basic information of genome stability related RNA binding proteins that immunoprecipitated from 293T cells by CCCH(5) domain of mouse ZC3H14.
|
Info. gi number |
Official Symbol |
Name |
Other Aliases |
Organism |
|
DHX9 |
DEAH (Asp-Glu-Ala-His) box polypeptide 9 |
DDX9, LKP, NDHII, RHA |
[Homo sapiens] |
|
|
AIFM2 |
apoptosis-inducing factor, mitochondrion-associated, 2 |
AMID, PRG3, RP11-367H5.2 |
[Homo sapiens] |
|
|
DDB1 |
damage-specific DNA binding protein 1, 127kDa |
DDBA, UV-DDB1, XAP1, XPCE, XPE, XPE-BF |
[Homo sapiens] |
|
|
HNRNPF |
heterogeneous nuclear ribonucleoprotein F |
HNRPF, MGC110997, OK/SW-cl.23, mcs94-1 |
[Homo sapiens] |
|
|
FXR1 |
fragile X mental retardation, autosomal homolog 1 |
FXR1P |
[Homo sapiens] |
|
|
HNRNPH1 |
heterogeneous nuclear ribonucleoprotein H1 |
DKFZp686A15170, HNRPH, HNRPH1, hnRNPH |
[Homo sapiens] |
|
|
HNRNPL |
Heterogeneous nuclear ribonucleoprotein L |
FLJ35509, HNRPL, P/OKcl.14, hnRNP-L |
[Homo sapiens] |
|
|
HNRNPR |
heterogeneous nuclear ribonucleoprotein R isoform 2 |
FLJ25714, HNRPR, hnRNP-R |
[Homo sapiens] |
|
|
PABPC1 |
Poly(A)-binding protein 1 |
PAB1, PABP, PABP1, PABPC2, PABPL1 |
[Homo sapiens] |
|
|
PABPC4 |
poly(A) binding protein, cytoplasmic 4 (inducible form) |
APP-1, APP1, FLJ43938, PABP4, iPABP |
[Homo sapiens] |
|
|
PABPN1 |
poly(A) binding protein, nuclear 1 |
OPMD, PAB2, PABP2 |
[Homo sapiens] |
|
|
RAD50 |
RAD50 homolog (S. cerevisiae) |
RAD50-2, hRad50 |
[Homo sapiens] |
|
|
RBM10 |
RNA binding motif protein 10 |
GPATC9, GPATCH9, KIAA0122, MGC1132, MGC997, ZRANB5 |
[Homo sapiens] |
|
|
YWHAZ |
tyrosine 3-monooxygenase/ tryptophan 5-monooxygenase activation protein, zeta polypeptide |
KCIP-1, MGC111427, MGC126532, MGC138156 |
[Homo sapiens] |
As an example, we chose the dramatically up-regulated and relatively well studied RBM10 protein to confirm its interaction with the CCCH(5) domain. To test the interaction between Zc3h14 and RBM10, 293T cells were transfected to transiently express FLAG-EGFP-tagged CCCH(5). Lysates from the transfectants were subjected to immunoprecipitation with anti-FLAG antibody. The precipitates were analyzed with anti RBM10. The results demonstrated binding of FLAG-EGFP CCCH(5) to RBM10 (Figure 6D). In the reciprocal experiment, analysis of anti-RBM10 immunoprecipitates with anti-FLAG confirmed the interaction (Figure 6E). These data demonstrated that the Zc3h14 can interacted with RBM10 through its CCCH(5) domain. Since many complexes of RNA binding proteins are usually assembled in an RNA-dependent manner [26], we treated the lysates with RNase A to test whether CCCH(5)-FLAG-EGFP interacted with RBM10 in an RNA-dependent manner. The results showed that after RNase A treatment, RBM10 could not be immunoprecipitated by the anti-FLAG antibody (data not shown). These data demonstrated that the CCCH(5) domain interacted with RBM10 in an RNA-dependent manner.
DISCUSSION
Human and mouse ZC3H14 proteins are orthologues of yeast Nab2. Nab2 play important roles in yeast mRNAs regulation and cell viability [11]. Human ZC3H14 binds poly(A) through its C-terminus CCCH(5) domain. The poly(A) biding activity of ZC3H14 protein suggest that it may be required for normal cell viability and proliferation. However, the physiological and pathological roles of mammalian ZC3H14 remain unknown. In this study, we characterized the mouse Zc3h14 gene. Expression pattern analysis (by RT-PCR) showed that mouse Zc3h14 gene was widely expressed. Two transcript isoforms of mouse Zc3h14 gene were found. These data suggested that ZC3H14 might play multiple roles in various tissues. We found that mouse ZC3H14 protein not only localized to the nucleus, but also formed multiple nuclear speckles. The poly(A) binding activity, nuclear localization suggested that mouse Zc3h14 protein might play important roles in regulating mRNA processing and/or stability. The nuclear speckle formation of Zc3h14 protein suggested it may play some unique roles in RNA and/or DNA regulation. Serial partial deletion mutation and point mutation analysis were designed for NLS mapping. We had successfully mapped the core NLS sequences of mouse Zc3h14 protein to the 292 RKRK sequences. The C-terminus CCCH(5) deleted Zc3h14 still localized to the nucleus. However, it did not form nuclear speckles. The CCCH(5) domain itself lost the nuclear specific targeting. These data demonstrated that the CCCH(5) zinc fingers domain was necessary but not sufficient for ZC3H14 nuclear speckle formation. The biochemistry nature of the nuclear speckles needs to be characterized in future to explain the unique roles of Zc3h14.
Dominant negative mutants, especially the CCCH(5) domain of mouse Zc3h14 protein, caused cell growth delay. Multiple morphological abnormalities, such as the long processes grown out, the nuclear heterogeneity, multiucleation, and nuclear fragmentation, were easily found. DNA damage was one of the common reasons for cell growth delay [23,24]. The phenotypes observed in the CCCH(5) expressing cells suggested that DNA/ genome may be severely damaged. Immunostaining of γ-H2AX, a widely used specific marker for double strand DNA breaks, showed that the expression level of γ-H2AX was up-regulated and multiple nuclear foci were formed. These data indicated that over-expression of the CCCH(5) domain of mouse ZC3H14 protein induced DNA damage. We also detected a group of DNA damage/repair related genes to confirm the DNA damage. These genes included Cdk2, Ddb1, Mdm2, p21, p53, Rad50. p53 played a central role in DNA damage-induced repair and cell apoptosis [27-30]. Mdm2 was a target gene of the p53 and formed an autoregulatory negative feedback loop with p53 [27]. p53 regulated cyclin-dependent kinase inhibitors p21 and p27. p21 and p27 played regulatory roles in S phase DNA replication and DNA damage repair [28]. CDK2 played a central role in DNA damage–induced cell cycle arrest and DNA repair [29]. DDB1 played an essential role in maintaining viability and genomic integrity of dividing cells [30]. RAD50 protein played important roles in cellular response to DSBs [31]. As expected, these genes were dramatically up-regulated. The RT-PCR results further indicated that DNA damage had been induced by over-expression of the CCCH(5) domain.
We explored the potential molecular mechanisms of how the DNA damage was induced by the expression of the CCCH(5) domain. Our immunoprecipitation and mass spectrometry analysis demonstrated that, a group of RNA binding proteins, such as DHX9, HNRNPF, HNRNPH1, HNRNPL, HNRNPR, PABPC1, PABPC4, PABPN1 and RBM10, interacted with the CCCH(5) domain of Zc3h14. Most of these proteins correlated with cell apoptosis and/or DNA damage. DHX9 was recently reported to be related to growth arrest and apoptosis [32]. HNRNPF and HNRNPH was reported that play important roles in modulating the alternative splicing of the apoptotic mediator Bcl-x [33]. More RNA-binding proteins, PABPC1, PABPC4, PABPN1 and RBM10 were also correlated with apoptosis and anti-virus response [34-36]. AIFM2 (AMID) was an p53-inducible, AIF-homologous and mitochondria-associated protein that has been implicated in caspase-independent apoptosis [37]. FXR1 was an autoantigen processed during apoptosis [38]. RAD50 was an structural maintenance of chromosomes (SMC) protein family member. It participated in a variety of cellular processes, including DNA double-strand break repair, cell cycle checkpoint activation and telomere maintenance [39]. YWHAZ (KCIP-1) functioned as a anti apoptosis protein. Inhibition its expression induced apoptosis [40]. As an example, we further showed that the CCCH(5) domain interacted with RNA binding protein RBM10, in an RNA dependent manner. Interactions between these genome stability and DNA damage related proteins and Zc3h14 CCCH(5) domain provide more supports to our novel finding that mammalian is a genome stability and DNA damage/repair associated protein.
One of the important questions to be answered is how ZC3H14 works in vivo. It was reported that ZC3H14 bound poly(A) through the CCCH(5) domain [12]. Poly(A) binding proteins (Pabs) play key roles in regulating posttranscriptional mRNA translation and degradation [4-6]. It was one of the possibilities that the CCCH(5) domain of Zc3h14 caused DNA damage through affecting functions of mRNPs. Then, some mRNAs that encoding chromatin stability related proteins were abnormally translated or degraded and finally, they affected genome/chromatin stability and caused DNA damage. More biochemistry studies need be done to find out the interaction network of mammalian ZC3H14 Conditional knockout mice will be very helpful to understand the functions of Zc3h14 during development and in multiple adult tissues.
ACKNOWLEDGEMENT
We appreciate Dr. Trono (NCCR, Switzerland) for providing the lentivirus vectors. We thank the Proteomics Core Facility at NIBS for performing the mass spectrometry analysis. This project was partially supported by National High Technology Project 863 (2005AA210930).