Evaluation of Next Generation ?lactam Inhibitors against Penicillinase Producing Methicillin-Resistant Staphylococcus aureus
- 1. Department of Genetics, Osmania University, India
- 2. Department of Zoology, Osmania University, India
- 3. Department of Botany, Osmania University, India
Abstract
Antibiotics are indispensable drugs in curing pathogenic bacteria that cause deadly diseases and infections. A plethora of antibiotics are available in different classes, however, βlactam inhibitors are one of the oldest class of inhibitors against manygram-positive as well as gram-negative bacteria. Pathogens are becoming more powerful, gradually developing resistance against successful βlactam antibiotics. Therefore, synthesis of novel βlactam inhibitors is a fundamental process in drug discovery. This study focused on the Methicillin-Resistant Staphylococcus aureus (MRSA) that produces penicillinase: PBP3. We identified 14 popular βlactam inhibitors: Ceftobiprole, Clavulanate, Sulbactam, Tazobactam, Biapenem, Razupenem, LK-157, Ceftolozane, Ampicillin, Amoxicillin, Ticarcillin, Piperacillin, Penicillin G, and Cefotaxime. Amongthem, Clavulanate, Sulbactam, and Tazobactam are next generation βlactam antibiotics. Molecular docking was carried out between MRSA-PBP3 and βlactam inhibitors in order to report the potential ligand. Interestingly, we found Clavulanate – a next generation antibiotic along with ther leading molecule in a row (Sulbactam, and Tazobactam) – has shown least binding affinity when compared with reference molecule Penicillin-G. Despite Clavulanate effectively acting against MRSA, its binding stability is less in MRSA-PBP3 complex. Therefore this study suggesting improved analogs are required to strengthen binding interaction alongside active against resistant bacteria.
Keywords
β-Lactam antibiotics , Penicillin-binding proteins (PBPs) , Molecular docking
Citation
Setti A, Sanat Kumar C, Naresh B, Someswar Rao S, Pawar SC (2017) Evaluation of Next Generation βlactam Inhibitors against Penicillinase Producing Methicillin-Resistant Staphylococcus aureus. JSM Biol 2(2): 1014.
INTRODUCTION
Since the discovery of antibiotic, it has been playing lavished role in the treatment of the bacterial diseases. In the past decade, usage of antibiotics is increasing several folds. Although bacteria could be killed by antibiotics, bacteria is gradually developing the antibiotic resistance due to long-term exposure to a spectrum of antibiotics. Across the board, hospitals are the sources of multidrug-resistant bacteria. The gram positive and negative; bacilli and cocci: Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter species, Pseudomonas aeruginosa, and Enterobacter species, reported as ESKAPE bacteria, develop antimicrobial resistance [1,2]. Perhaps, comprehensive chemotherapy and immunotherapy are required to steer the infectious bacteria [3]. Besides, there is a growing demand for novel antibiotic discovery to meet the irreproachable control of the antimicrobial resistant microbes [4]. Although penicillin, a β-lactam antibiotic, is first efficient antibiotic drug to be used to treat previously serious Staphylococcus and Streptococcus infections, now no longer successful [5,6]. After penicillin, next generation drug, methicillin is also a class of narrow-spectrum penicillin antibiotic that is used to treat penicillinase-producing bacteria Staphylococcus aureus. However, certain S. aureuss trains are resistant to methicillin; called methicillin-resistant Staphylococcus aureus (MRSA). In addition, in certain populations, community-associated MRSA cause skin and soft tissue infections in India and other countries [7-9]. Therefore, combination antibiotic treatment for MRSA has been adopted
In gram-positive bacteria thick peptidoglycan layer, while in gram-negative thin peptidoglycan layer, contain N-acetylglucosamine-N-acetyl muramic repeats. The cell membrane of the bacteria contains penicillin-binding proteins (PBPs), a family of acyl-serine transferases, involved in the synthesis of the peptidoglycan layer of the cell wall [10]. PBPs are classified into two categories based on their molecular weight: indispensable high molecular weight (HMW) group and dispensable low molecular weight (LMW) group. HMW group PBPs are further divided into class A and class B [11]. Former involves in glycosyltransferase and transpeptidase, while later involves only in transpeptidase activity. Covalent and noncovalent binding energies between Penicillin and PBPs were studied by CD and fluorescence experiments [12]. New generation β lactamases are always sought to enhance the drug efficacy to curb the βlactam resistant bacteria. Consequently, an insilico high throughput screening for novel molecules against PBPs of Staphylococcus aureus and Neisseria gonorrhoeae were sought for anti-gonococcal agents [13,14]. Likewise, molecular docking and dynamics studies were also reported on PBP inhibitors – it has been found in molecular dynamics simulations that Ceftobiprole bound PBP complex is more stable than free PBP [15]. In addition to this, several anti microbial targets are being investigated in insilico approach to confront the pathogenic bacteria. Therefore, in the search of next generation PBP inhibitors, we identified 14 compounds to evaluate their efficacy on PBP.
METHODOLOGY
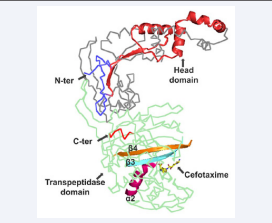
This study mainly focused on MRSA that contain HMW, class B, PBP3. The crystal structure of this protein was obtained from Protein Data Bank (PDB) database with 3VSL as PDB identifier [16]. Two chains of homodimer structure of 3VSL were separated in Swiss PDB Viewer (SPDBV) tool; Chain A possesses βlactam drug, cefotaxime, in the activesite. PBP3 of MRSA comprises 691 residues – UniProt id: Q8NWC2, however, crystal structure available for 633 residues: ALA-46 to Lys-678. Two domains were observed in MRSA-PBP3: N-terminal head domain and C-terminal transpeptidation domain as shown in Figure (1). In protein-ligand interaction, active site key residues participate in seldom irreversible covalent, or most often reversible noncovalent interactions. PDB structure of 3VSL, MRSA-PBP3, is cocrystallized with Cefotaxime; SER-392 is identified as active site residue that is, infact, formed a covalent bond with the ligand.
Figure 1 Crystal structure of Penicillin-Binding Protein 3 (PBP3) of Methicillin-resistantbacteria Staphylococcus aureus (MRSA), PDB id: 3VSL. PBP3 has Head domain at N-terminal and Transpeptidase domain at C-terminal. Cefotaxime is shown in the active site of the protein-ligand complex.
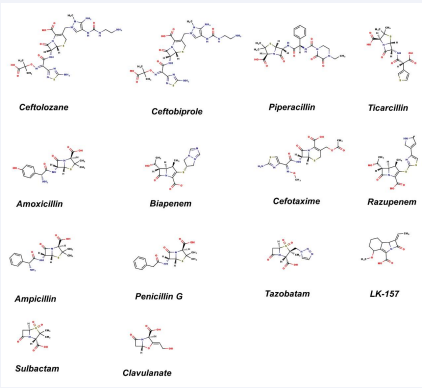
The goal of the study is to report the better β lactam inhibitor; validate the next generation inhibitors of the PBPs. Therefore in the present study; 14 ligands: Ceftobiprole, Clavulanate, Sulbactam, Tazobactam, Biapenem, Razupenem, LK-157, Ceftolozane, Ampicillin, Amoxicillin, Ticarcillin, Piperacillin, Penicillin G, and Cefotaxime (Figure 2); those drugs which are approved and need to be approved have been considered for molecular docking. In the beginning of 2014, it was introduced in a review that some of the β lactam antibiotics have been restored with Clavulanic Acid, Sulbactam, and Tazobactam. In the same review; Biapenem, Razupenem, LK-157, and Ceftolozane were also reported which are yet to be approved [17]. Three-dimensional structures of the ligands with Explicit Hydrogens were sketched and geometry was cleaned in Marvin Sketch tool.
Figure 2 β lactam inhibitors of Penicillin Binding Protein 3 (PBP3) used for molecular docking to endow the better interacting molecule.
Molecular docking is an inexpensive technique to determine the binding interaction stability of the compound, this technique is indispensable in drug discovery process. However, further experimental evidence is required to correlate the insilico and invitro data. Identified 14 β-lactam inhibitors docked against MRSA-PBP3 in Molegro Virtual Docker (MVD) tool. This tool reproduces binding energy/ free energy of the docked complex in the form of Mol Dock Score Grid (MDS-Grid), Mol Dock Score (MDS), and Rerank Score (RRS). These scores aptly explain the stability of the ligand molecule in the docked protein-ligand complex. In this docking experiment, following parameters were used in the docking wizard: MDS-Grid scoring function, 0.2 Å Grid resolution, and default search algorithm MolDock SE. During docking 15 Å search space/grid for the ligand is set in a spherical format with 16.55, -48.00, and 25.65Å as sphere center coordinates that retain the active site residue SER-392. Prior to setting docking experiments, MVD tool had been validated for accuracy by docking native ligand Cefotaxime against MRSAPBP3. After three redocking attempts, root mean square deviation (RMSD) between the crystal and sketched Cefotaxime is found to be 1.02 Å. Furthermore, docked complexes were visualized in PyMOL
RESULTS
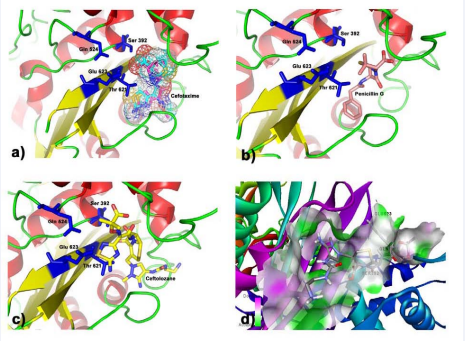
Figure 3 Activesite of MRSA-PBP3 with ligand: a) Validation of MVD with Cefotaxime. RMSD between crystal pose and docked pose is 1.02 Å; b) Penicillin G, reference molecule, is shown in the active site of PBP3 to compare the docking scores of the other beta-lactam inhibitors; c) Ceflolozane, top-ranked molecule in the docking study, is shown in the active site of PBP3 along with its key residues; d) H-bond donor (Pink) and acceptor (Green) of cavity is seen around the top ranked molecule Ceflozane.
MVD docking program was used in order to anticipate the binding stability of the 14 ligands: Ceftobiprole, Clavulanate, Sulbactam, Tazobactam, Biapenem, Razupenem, LK-157, Ceftolozane, Ampicillin, Amoxicillin, Ticarcillin, Piperacillin, Penicillin G, and Cefotaxime. In fact, in the β lactam kind of drugs, Penicillin G was survived for a long time in the market battle against gram-positive bacteria. Thus, in the present study, Penicillin G and MRSA-PBP3 docked complex’s MDS-Grid has been served as a reference to compare with a remaining set of ligands. Crystal structure of the MRSA-PBP3, PDB ID:3VSL, is used for protein target for identified set of 13 ligands. The active site is located in the C-terminal region; α2 helix, β3 and β4 strands form binding interactions with Cefotaxime. Predicted volume and surface area of the binding pocket is 256 Å3 and 782 Å2 . Active site residues SER-392, GLN-524, THR-621, and GLU-623 are involved in H-bond formation with the ligand. Cefotaxime is present in the crystal structure of the MRSA-PBP3 (PDB id: 3VSL); to check the functional status of the MVD tool we sketched the ligand in Marvin sketch. The 3D geometry of the sketched molecule with explicit hydrogens had been optimized and docked over the PBP3. Likewise, remaining molecular structures were also sketched and optimized after adding explicit hydrogens. Molecular docking was accomplished in Molegro Virtual Docker (MVD) tool, mol dock score – Grid (MDS-Grid), mol dock score (MDS) and rerank score, root mean square deviation (RMSD), and torsions for each ligand is shown in Table 1. In docking, noncovalent interactions H-bond donors, H-bond acceptors play a crucial role in stability; Ceftolozane formed H-bonds with Ser-392, Lys-395, Lys-427, Ser-429, Ser-448,Lys-600,Gly-602,Thr-603, Leu-663 as shown in Figure 3; while Penicillin G (reference molecule) formed with His-447, Ser-448, Thr-621.
Table 1: Penicillin Binding Protein 3 (PBP3) of Methicillin Resistance Staphylococcus aureus (MRSA) used as atarget to dock β lactam inhibitors in Molegro Virtual Docker (MVD) tool. Cefotolozane has shown strong interaction with MRSA-PBP3.
| S.No. | Ligand | MDS-Grid | MDS | Rerank Score | RMSD | Torsions | Docking attempts |
|---|---|---|---|---|---|---|---|
| 1 | Ceftolozane | -220.37 | -185.18 | -128.91 | 0.89 | 14 | 8 |
| 2 | Ceftobiprole | -186.49 | -162.21 | -104.88 | 0.53 | 6 | 2 |
| 3 | Piperacillin | -183.68 | -162.60 | -97.09 | 0.71 | 8 | 5 |
| 4 | Ticarcillin | -163.33 | -132.70 | -73.16 | 1.23 | 4 | 2 |
| 5 | Amoxicillin | -159.73 | -136.78 | -99.92 | 1.73 | 4 | 4 |
| 6 | Biapenem | -158.40 | -139.53 | -66.22 | 0.80 | 8 | 2 |
| 7 | Cefotaxime | -152.69 | -135.66 | -100.80 | 1.02 | 4 | 3 |
| 8 | Razupenem | -144.05 | -132.36 | -98.51 | 1.42 | 4 | 9 |
| 9 | Ampicillin | -138.69 | -128.04 | -87.41 | 0.83 | 4 | 3 |
| 10 | PenicillinG | -131.61 | -128.41 | -90.69 | 0.03 | 2 | 3 |
| 11 | Tazobatam | -128.50 | -100.50 | -72.92 | 0.45 | 2 | 6 |
| 12 | LK-157 | -111.71 | -91.32 | -36.66 | 0.01 | 1 | 3 |
| 13 | Sulbactam | -105.48 | -84.92 | -66.51 | 0.91 | 2 | 4 |
| 14 | Clavulanate | -103.35 | -77.27 | -62.90 | 0.43 | 5 | 6 |
DISCUSSION
Cefotolozane has shown strong binding interaction with PBP3 because it comprised of 75 atoms with 14 torsional bonds which let the molecule so flexible that allows 18 H-bonds. Interestingly, each one of O(8), and O(2) atom of the Cefotolozane formed 3 H-bonds. Furthermore, the orientation of this ligand tightly fitted in the pocket of the PbP3 (Figure 3c and 3d) that sets the protein to be non-functional. Apparently, Penicillin G was very effective βlactamantibiotic since its discovery, however, its efficacy detained due to developed resistance in bacterial strains. Therefore, it is considered to be a reference molecule to compare the next generation βlactam antibiotics. In fact, Penicillin G in the PBP3 binding pocket formed 3 H-bonds, while Cefotolozane formed 18 H-bonds. A short while ago, it was pointed in a review that Clavulanate, Sulbactam, and Tazobactam are sort of next generation βlactam antibiotics which may replace the existing Ampicillin, Amoxicillin Ticarcillin, and Piperacillin, but in this study we find they show less efficacy than Penicillin G – a reference molecule to compare binding affinity. In conclusion, Cefotolozane is most effective; Clavulanate is least effective against MRSA-PBP3. Clavulanate, Sulbactam, and Tazobactam probably effective against another kind of PBPs but not for MRSAPBP3, this observation suggests most of the antibiotics are not effective for all kind of bacteria. It is also not escaped our notice the next generation promising molecules Clavulanate, Sulbactam, Tazobactam and LK-157 analogs perhaps improves the binding interactions with MRSA-PBP3.
CONCLUSION
Penicillin-binding proteins are potential targets for a wide variety of pathogenic gram-positive bacteria. Since the discovery of penicillin several pathogenic bacteria including Staphylococcus aureus, Neisseria meningitides, and Listeria monocytogenes were controlled. However, over the period of time, the bacteria gradually developed the resistance against βlactam kind of inhibitors. Thus, its successors Methicillin, Ampicillin, Flucloxacillin, and Dicloxacillin were released into the market. Once a powerful drug Methicillin was ineffective against penicillinase or βlactam group degrading bacteria S.aureus. In the present study, 14 popular βlactam inhibitors were collected and docked against MRSA-PBP3 in order to check the potency of the each candidate βlactam inhibitor. Interestingly, Ceftolozane has shown strong binding interaction while Clavulanate– a next generation βlactam inhibitor – has shown least binding interaction with PBP3. Although Clavulanate produced aleast binding score in molecular docking, it probably controls the MRSA compared with Ceftolozane. Finally, this study points out the essence of the improvement of next generation antibiotics in view of improving binding strength and controlling penicillinaseproducing bacteria by competing for the resistance.
ACKNOWLEDGEMENT
The authors would like to thank the Head of the Department, Genetics for providing necessary facilities to carry out the work. Author ARS would like to thank UGC for funding the research work.
REFERENCES
7. Uzunovic S, Bedenic B, Budimir A, Ibrahimagic A, Kamberovic F, Fiolic Z, et al. Methicillin-resistant S. aureus (MRSA), extended-spectrum (ESBL)- and plasmid-mediated AmpC ss-lactamase -producing Gramnegative bacteria associated with skin and soft tissue infections in hospital and community settings. Medicinski glasnik : official publication of the Medical Association of Zenica-Doboj Canton, Bosnia and Herzegovina. 2015; 12: 157-168.