What are the Genomunities that Govern Sexual Dimorphisms?
- 1. Department of Animal Sciences and Center for Reproductive Biology, Washington State University, USA
- 2. Animal Breeding and Genetic Unit, University of New England, Australia
Abstract
Here we propose a new concept called genomunities (genome communities) and thus explore how they govern 116 sexual dimorphism pathways in Xenopus tropicalis. Based on alternative polyadenylation events, a total of 262 genomunities were identified to be associated with these pathways. These genomunities originated from six sources: oocytes (maternal RNA), oocytes/zygotes, zygotes, embryos, adult females and adult males. Our discussion is focused on genomunity memberships, genomunity contribution and genomunity partnerships. Overall, our genomunity concept may open new research fronts to understand emerging biological and biomedical questions related to gender and gender expression, sex and gender identity, transgender, gender dysphoria, and sexual orientation, for example.
Keywords
Genomunity, Membership, Contribution, Partnership, Sexual dimorphisms
Citation
Jiang Z, Carrion SA, Michal JJ, Zhou X, Wang H, et al. (2022) What are the Genomunities that Govern Sexual Dimorphisms? JSM Sexual Med 6(3): 1089.
INTRODUCTION
The term “sexual dimorphism” is used to describe the dramatic differences in phenomes between males and females of the same species. Primarily, males and females differ in reproductive systems. In mammals, for instance, the vagina, uterus, fallopian tubes, vulva and ovaries are female-specific reproductive organs [1], while the penis, scrotum, testes, epididymis, vas deferens, prostate and seminal vesicles are male-specific reproductive systems [2]. Furthermore, males and females each have dominant hormonal profiles that initiate, form, and develop unique reproductive systems and ensure each function appropriately [1,2].
At the genome level, on the other hand, males and females seem equal. In mammals, for example, males and females share the same autosomes, but differ in sex chromosomes. Females have two X chromosomes, while males have only one. The Y chromosome is male-specific. In comparison to the X chromosome, relatively few genes reside on the Y chromosome. However, genes on one of the X chromosomes are silenced due to a process called X inactivation. Interestingly, 15% - 30% of the genes on the inactivated X chromosome remain active [3]. With the mechanisms, the dosage differences are actually compromised for balance between sexes.
In addition to reproductive systems, no doubt, gender does affect many physical, physiological, pathological and psychological processes, such as adaptation to space, blood pressure control, body fluid homeostasis, circadian rhythms, diseases and aging, exercise endurance, pain and its relief, pitch range and speech fundamental frequency, skeletal muscle kinetics and substance use addiction [4-13], for example. Even the current Covid19 pandemic is biased against men as they have stronger response to the virus, greater morbidity and higher mortality than women [14,15]. Therefore, how structurally equal genomes result in so many phenotypic differences between genders remain mysterious.
Interestingly, almost every gene uses alternative promoters, exons and/or polyadenylation sites to maximize genome functions [16]. In fact, timely use of transcript variants is essential to maintain differentiation and development, and respond appropriately to environmental challenges, while misuse of alternative transcripts often causes defects, diseases and disorders [17-19]. By examination of over 18,000 proteincoding genes in 23 cell types from 798 human tissue samples, Reyes and Huber [20] discovered that transcriptome diversity was largely due to usage of alternative transcription start (ATS) and polyadenylation (APA) sites, rather than by use of alternative splicing. As such, we have developed both whole transcriptome start and termini site sequencing (WTSS-seq and WTTS-seq) methods to profile RNA diversity [21,22].
In addition to mammals, sexual dimorphisms have been well observed in amphibians [23,24]. By profiling of APA events using Xenopus tropicalis embryos and whole bodies of adults, in particular, we found 116 pathways related to sexual dimorphisms in the species (Supplemental Table 1, [22]). For APA sites derived from embryos at stages 6, 8, 11, 15 and 28, we further classified them into origins of oocytes, oocytes/zygotes and zygotes. Any APA sites that did not clearly fit into these three categories were classified as embryonic origin. In addition, we have information on both ATS and APA events because both adult males and females were also used in the study [22]. As these RNA variants have different origins executed to complete their functions at different stages, here we would coin a new term “genomunity” (genome community) to illustrate how they are coordinated to govern the sexual dimorphisms.
Sexual dimorphisms – single vs. multiple genomunities
As shown in Supplemental Table 1, forty-six of the 116 pathways associated with sexual dimorphisms are each supported by a single genomunity. The remaining 70 sexual dimorphism processes were the result of the coordination of more than one genomunity. These multiple genomunities underlie 33, 17, 5, 11 and 4 pathways with APA-associated genes produced by either two, three, four, five or six origins, respectively. As such, these 116 sexual dimorphisms are coordinated by a total of 262 genomunities (Supplemental Table 1). The size of a genomunity varies, ranging from 3 genes to 315 genes per pathway. Overall, genomunities are variable in numbers of genes and their contents.
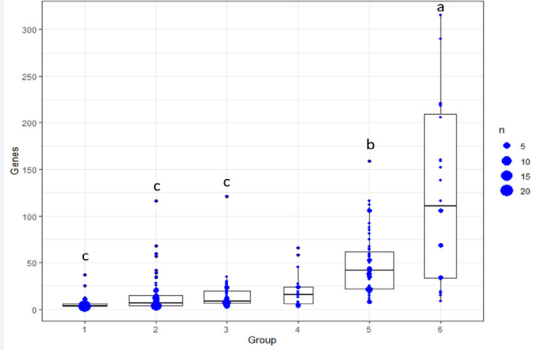
Figure 1: Number of genes per genomunity. It depends on the numbers of genomunities involved in a specific sexual dimorphism pathway. 1 – 6 indicate one genomunity to six genomnities per pathway.
For example, the single-origin genomunities have the lowest numbers of genes with an average of 5.74 per pathway (Figure 1). The averages increase to 14.38, 14.68 and 19.45 genes per pathway for two-, three- and four-source genomunities, respectively. The average number of genes per pathway dramatically jump to 49.33 and 122.58 for sexual dimorphism processes involving five and six sources of genomunities (P<0.0001). Furthermore, the number of members per genomunity varies a lot, with a range of 3 – 37, 3 – 116, 3 – 121, 3 – 66, 8 – 159, and 9 – 315 genes for each of these six categories (Supplemental Table 1). Therefore, the more complex the sexual dimorphism is, the more genomunities it needs with higher numbers of genes per genomunity to complete the complex process (Supplemental Table 1).
In addition to the multiple genomunities for single sexual dimorphism pathway, there are also 20 cases that appear to possess a single genomunity for multiple sexual dimorphism pathways (Supplemental Table 1). These genomunities each have 3 to 69 genes, responsible for two to four sexual dimorphism pathways. Among these 20 cases, 16 are also included in multiple genomunities to support different reproductive functions. In addition, only three genomunities each also have different origins (Supplemental Table 1). No doubt, there are always genomunities with similar contents of genes functioning on different pathways. As one gene often executes alternative transcript events, here we would argue that sometimes, the same genes may or may not correspond to the same APA events [22].
Sexual dimorphisms – before vs. after birth genomunities
These 116 sexual dimorphism pathways governed by these 262 genomunities can be further classified into seven biological clusters (Supplemental Table 1). The gonad, germ cell and gamete cluster possess 67 genomunities, mainly dealing with development, differentiation, division, generation, migration, and/or proliferation of gametes, germ cells, Leydig cells, oocytes, Sertoli cell, sperm and gonad. The reproductive hormone cluster includes processes related to estradiol, estrogen, estrogen receptor signaling, follicle-stimulating hormone, gonadotropin, progesterone and progesterone receptor signaling with a total of 31 genomunities. The mammary gland and lactation cluster involves 41 genomunities, promoting development, differentiation, formation, involution, morphogenesis and/or proliferation of mammary gland and its alveolus, duct, epithelial cell, epithelium, lobule and placode. The smallest cluster is related to reproductive organs, such as genitalia and vagina with 9 genomunities. Next is a 30-genomunity cluster that focuses on development, growth and morphogenesis and/or regulation of prostate gland and its epithelium and stromal, prostate glandular acinus and prostatic bud. The reproductive process cluster owns the second largest number of 53 genomunities, regulating estrous cycle, pregnancy, fertilization, maternal behavior, mating, ovulation and its cycle, reproductive process and sex determination and differentiation. Lastly, the reproductive system cluster holds a total of 31 genomunities related to reproduction in multicellular organism, reproductive structure and systems, and sexual characteristics.
In many organisms, a life starts with a fertilized egg or oocyte, which contains RNA molecules provided by the mother, or so-called maternal RNA. In early embryo development, there is a period called the maternal-to-zygotic transition. During this period, maternal RNA is degraded, allowing the zygote to take over the control of its development [25]. Therefore, the APA genomunities have three clear origins: oocytes, oocytes/ zygotes and zygotes. As defined by us previously, the oocytegenomunities are those APA sites with expression terminated at stages 8, 11, 15 or 28 [22]. The APA events that are defined as oocyte/zygote origin are those that are expressed initially in oocytes, but are reused by the zygotes following degradation of the maternal copies. The zygote-origin APA sites are those that are silent until they are expressed at stages 11, 15 or 28 [22]. The remaining APA events that appeared before birth, but are absent in oocytes, oocytes/zygotes and zygote sources are collectively called the embryo origin. Nevertheless, oocytes, oocytes/zygotes, zygotes and embryos provide all before-birth genomunites. Consequently, all after-birth genomunities are derived from adult males and adult females.
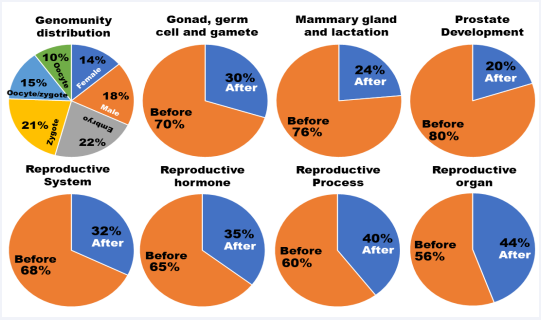
Figure 2: Genomunity contribution. The before-birth genomunites are provided by oocytes, oocytes/zygotes, zygotes and embryos, while the after-birth genomunities are produced by adult males and adult females.
As shown in Figure 2 (genomunity distribution), all 262 genomunities are distributed as 68% (oocytes – 10%, oocytes/ zygotes – 15%, zygotes – 21% and embryos – 22%) for the beforebirth and 32% (females – 14% and males – 18%) for the afterbirth categories. Interestingly, the sexual dimorphism pathways grouped in the reproductive systems cluster follow the same distribution. However, the before-birth contribution increases from 68% to 70% for the gonad, germ cell and gamete cluster, 76% for the mammary gland and lactation cluster and even 80% for the prostate and its related event cluster. These results imply that the foundation of sexual dimorphisms is mainly built during the before-birth period. On the other hand, the after-birth shares also increase from 32% to 35% for the reproductive hormone cluster, 40% for the reproductive process cluster and 44% for the reproductive organ clusters. It seems that the after-birth period plays an important role in regulating reproductive cycles, mating behaviors and sexual appearance. Overall, the before- and afterbirth contributions significantly differ among these clusters (χ2 = 16.726, P = 0.01034478).
Sexual dimorphisms – male vs. female genomunities
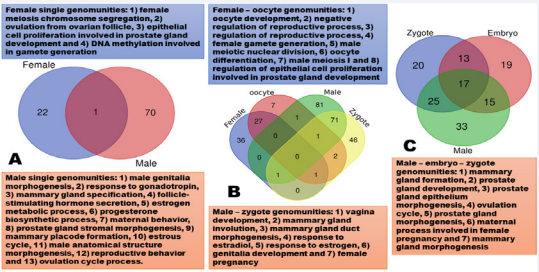
As demonstrated in Figure 3A and Supplemental Table 1, adult males alone provide APA-associated genes to form 13 single genomunities, while the female’s contrition is only four single genomunities. On basis of their underlying genes, only one is overlapped between females and males’ contribution. For sexual dimorphisms governed by two genomunities, adult males dominantly partner with zygotes, while the adult females and oocytes are the most favorable collaborators to guide their unique pathways. As shown in Figure 3B, the former partnership jointly supports seven sexual dimorphism pathways, but the latter combination regulates eight pathways. Interestingly, the partner combination does not differ significantly (P > 0.05) in gene sharing rate: 52.33% for the male – zygote pair vs. 53.85% for the female – oocyte pair.
Figure 3: Genomunity partnerships. A. single genomunities; B. two-genomunity partnerships; and C. three-genomunity partnerships.
For sexual dimorphisms requiring involvement of three genomunities, the most dominant one is the combination of adult males with both zygotes and embryos, which is associated with seven sexual dimorphism pathways (Figure 3C). As shown in Supplemental Table 1, the second most dominant team is still related to the adult males, but paired with both oocyte/ zygote and zygotes, which guides 1) mammary gland epithelial cell proliferation, 2) mammary gland alveolus development and 3) mammary gland lobule development. Except the oocytes described above, unfortunately, adult females seem reluctant to extend their partnership further for collaboration with other sources of genomunities (Supplemental Table 1).
Here is a list of male pathways, but governed by the female genomunities. Somehow the female genome can execute alternative events to organize 1) male gamete generation, 2) male gonad development, 3) male meiosis I, 4) male meiotic nuclear division, 5) spermatid differentiation, 6) spermatogenesis, 7) epithelial cell proliferation involved in prostate gland development, 8) regulation of epithelial cell proliferation involved in prostate gland development, 9) male sex differentiation, and 10) development of primary male sexual characteristics (Supplemental Table 1).
On the other hand, males also do things that should be done by females, for example, such as 1) lactation, 2) vagina development, 3) estrous cycle, 4) female pregnancy, 5) maternal process involved in female pregnancy, 6) ovulation, 7) ovulation cycle process, 8) ovulation cycle, and 9) female sex differentiation (Supplemental Table 1). In Xenopus tropicalis, in fact, males have three types of sex chromosome combinations: YZ, YW and ZZ, while females possess either ZW or WW sex chromosomes [25]. Therefore, whether these genomunities are cofounded with the sex determination systems in the species warrants to be investigated further. Furthermore, the genetic improvement practice in dairy production has confirmed that selection of sires (males) can improve daughter pregnancy rate, gestation length, calving ability, and conception rate [26-28]. These results clearly indicate that males contribute significantly to the improvement of fertility and its related phenotypes in females.
SUMMARY
Using 116 sexual dimorphism pathways detected in Xenopus tropicalis embryos and adults as an example, we explore a new concept called genomunities and thus understand their membership, contribution and partnership rules in gender divergence and dynamics. When a biological pathway is controlled by a single genomunity, it usually needs relatively few members function. A complex phenotype, however, usually requires multiple genomunities, so each of them should have a large number of genes to work together. So, the basic trend is: the more complex the trait, the more genomunities are required with more genes included in each genomunity. In terms of genomunity – sexual dimorphism relationships, both one genomunity for multiple pathways and multiple genomunities for one pathway exist. Genomunity contribution largely relies on types of sexual dimorphisms. Prostate-, mammary gland- and gonad/germ cell/gamete-related events are largely coordinated by genomunities provided by oocytes, oocytes/zygotes, zygotes and embryos. For some events related to reproductive hormones, reproductive processes and reproductive organs, they need more genomunity events after birth. Males and females differ in choices of genomunity partnerships. The first choice for males to collaborate is zygotes, followed by embryos and oocytes/zygotes. The oocyte-produced genomuities are often aligned with those generated by females to regulate a phenotype. We believe that our genomunity concept can help understand physical, physiological, pathological and psychological mechanisms underlying gender and gender expression, sex and gender identity, transgender, gender dysphoria, and sexual orientation, for example.
ACKNOWLEDGEMENTS
This work was supported by the Eunice Kennedy Shriver National Institute of Child Health & Human Development of the National Institutes of Health under Award Number R21HD076845 and the National Institute of Food and Agriculture, United States Department of Agriculture under Award Numbers 2016-67015-24470, 2020-67015-31733 and 2022-51300- 38058 to ZJ. The research was also supported by funds provided for medical and biological research by the State of Washington Initiative Measure No. 171 and the Washington State University Agricultural Experiment Station (Hatch funds 1014918) received from the National Institutes for Food and Agriculture, United States Department of Agriculture to ZJ. Special thanks go to the Fulbright Australia for supporting ZJ’s stay at University of New England in 2022.
REFERENCES
1. Rosner J, Samardzic T, Sarao MS. Physiology, Female Reproduction. [Updated 2021 Oct 9]. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2022.
2. Gurung P, Yetiskul E, Jialal I. Physiology, Male Reproductive System. [Updated 2021 May 9]. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2022.
3. Balaton BP, Fornes O, Wasserman WW, Brown CJ. Cross-species examination of X-chromosome inactivation highlights domains of escape from silencing. Epigenetics Chromatin. 2021; 14: 12.
4. Harm DL, Jennings RT, Meck JV, Powell MR, Putcha L, Sams CP, et al. Invited review: gender issues related to spaceflight: a NASA perspective. J Appl Physiol (1985). 2001; 91: 2374-83.
5. Tadic M, Cuspidi C, Grassi G, Ivanovic B. Gender-specific therapeutic approach in arterial hypertension - Challenges ahead. Pharmacol Res. 2019; 141: 181-188.
6. Vivas L, Dadam FM, Caeiro XE. Sex differences in body fluid homeostasis: Sex chromosome complement influences on bradycardic baroreflex response and sodium depletion induced neural activity. Physiol Behav. 2015; 152(Pt B): 416-21.
7. Nicolaides NC, Chrousos GP. Sex differences in circadian endocrine rhythms: Clinical implications. Eur J Neurosci. 2020; 52: 2575-2585.
8. Sampathkumar NK, Bravo JI, Chen Y, Danthi PS, Donahue EK, Lai RW, et al. Widespread sex dimorphism in aging and age-related diseases. Hum Genet. 2020; 139: 333-356.
9. Tiller NB, Elliott-Sale KJ, Knechtle B, Wilson PB, Roberts JD, Millet GY. Do Sex Differences in Physiology Confer a Female Advantage in UltraEndurance Sport? Sports Med. 2021; 51: 895-915.
10.Pieretti S, Di Giannuario A, Di Giovannandrea R, Marzoli F, Piccaro G, Minosi P, Aloisi AM. Gender differences in pain and its relief. Ann Ist Super Sanita. 2016; 52: 184-9.
11.Konomi U, Watanabe Y, Komazawa D. Sex Differences in Pitch Range and Speech Fundamental Frequency After Arytenoid Adduction and Thyroplasty. J Voice. 2016; 30: 362-70.
12.Haizlip KM, Harrison BC, Leinwand LA. Sex-based differences in skeletal muscle kinetics and fiber-type composition. Physiology (Bethesda). 2015; 30: 30-9.
13.Harp SJ, Martini M, Lynch WJ, Rissman EF. Sexual Differentiation and Substance Use: A Mini-Review. Endocrinology. 2020; 161: bqaa129.
14.Alwani M, Yassin A, Al-Zoubi RM, Aboumarzouk OM, Nettleship J, Kelly D, Al-Qudimat AR, Shabsigh R. Sex-based differences in severity and mortality in COVID-19. Rev Med Virol. 2021.
15.Wray S, Arrowsmith S. The Physiological Mechanisms of the SexBased Difference in Outcomes of COVID19 Infection. Front Physiol. 2021; 12: 627260.
16.Jiang Z, Zhou X, Li R, Michal JJ, Zhang S, Dodson MV, et al. Whole transcriptome analysis with sequencing: methods, challenges and potential solutions. Cell Mol Life Sci. 2015; 72: 3425-39.
17.López-Urrutia E, Campos-Parra A, Herrera LA, Pérez-Plasencia C. Alternative splicing regulation in tumor necrosis factor-mediated inflammation. Oncol Lett. 2017; 14: 5114-5120. 18.Martínez-Montiel N, Anaya-Ruiz M, Pérez-Santos M, MartínezContreras RD. Alternative Splicing in Breast Cancer and the Potential Development of Therapeutic Tools. Genes (Basel). 2017; 8: E217.
19.Reble E, Dineen A, Barr CL. The contribution of alternative splicing to genetic risk for psychiatric disorders. Genes Brain Behav. 2018; 17: e12430.
20.Reyes A, Huber W. Alternative start and termination sites of transcription drive most transcript isoform differences across human tissues. Nucleic Acids Res. 2018; 46: 582-592.
21.Zhou X, Li R, Michal JJ, Wu XL, Liu Z, Zhao H, et al. Accurate Profiling of Gene Expression and Alternative Polyadenylation with Whole Transcriptome Termini Site Sequencing (WTTS-Seq). Genetics. 2016; 203: 683-97.
22.Zhou X, Zhang Y, Michal JJ, Qu L, Zhang S, Wildung MR, et al. Alternative polyadenylation coordinates embryonic development, sexual dimorphism and longitudinal growth in Xenopus tropicalis. Cell Mol Life Sci. 2019; 76: 2185-2198.
23.Maan ME, Cummings ME. Sexual dimorphism and directional sexual selection on aposematic signals in a poison frog. Proc Natl Acad Sci U S A. 2009; 106: 19072-7.
24.Tadros W, Lipshitz HD. The maternal-to-zygotic transition: a play in two acts. Development. 2009; 136: 3033-42.
25.Roco ÁS, Olmstead AW, Degitz SJ, Amano T, Zimmerman LB, Bullejos M. Coexistence of Y, W, and Z sex chromosomes in Xenopus tropicalis. Proc Natl Acad Sci U S A. 2015; 112: E4752-61.
26.Lima FS, Silvestre FT, Peñagaricano F, Thatcher WW. Early genomic prediction of daughter pregnancy rate is associated with improved reproductive performance in Holstein dairy cows. J Dairy Sci. 2020; 103: 3312-3324.
27.Chebel RC, Veronese A. Associations between genomic merit for daughter pregnancy rate of Holstein cows and metabolites postpartum and estrus characteristics. J Dairy Sci. 2020; 103: 10754-10768.
28.Fang L, Jiang J, Li B, Zhou Y, Freebern E, Vanraden PM, et al. Genetic and epigenetic architecture of paternal origin contribute to gestation length in cattle. Commun Biol. 2019; 2: 100.