Haplotype Analysis of Heat Shock Protein70 Gene and Their Association with Essential Hypertension
- 1. Dr. B. R. Ambedkar Center for Biomedical Research, University of Delhi, India
- 2. Department of Cardiology, All India Institute of Medical Sciences, India
- 3. Department of Pharmacology, All India Institute of Medical Sciences, India
Abstract
Background: Essential hypertension affects major parts of worldwide population and is a potential risk factor for coronary artery disease and myocardial infarction, heart failure, stroke and renal failure. The present study aims to investigate the involvement of gene variants of HS70/1, HSP70/2 and HSP70/hom in essential hypertension in Northern Indian population.
Methods: 280 patients with essential hypertension and 261 healthy controls were recruited for the investigation. Genotyping of HSP70 polymorphisms (HSP70/1: +190G/C (rs1043618), HSP70/2:+1267A/G (rs1061581), and HSP7/hom: +2437T/C (rs2227956), were identified by PCR-RFLP techniques.
Results: A statistically significant difference in genotype distribution was found in HSP70/1 +190G/C [χ2 = 5.84, p = 0.015, Odds ratio = 1.53 (1.08-2.17) at 95% CI] and HSP70/hom +2437T/C [χ2 = 7.46, p = 0.006, Odds ratio = 1.60 (1.14-2.25) at 95% CI] polymorphisms in patients with essential hypertension in comparison to control group. Haplotype analysis indicated a significant r2 value (0.8) for HSP70/1 (rs1043618) and HSP70/2 (rs1061581) genotypes in patient group, whereas HSP70/ hom (rs2227956) stands independently and do not shows any association with other polymorphisms.
Conclusions: Our findings indicate that functional polymorphisms in HSP70 gene and their interaction are associated with the risk of essential hypertension in North Indian subjects.
Keywords
• Heat shock protein70
• Single nucleotide polymorphisms
• Essential hypertension
• Haplotype interaction
• PCR-RFLP
Citation
Srivastava K, Chauhan S, Chandra S, Narang R, Bhatia J, et al. (2016) Haplotype Analysis of Heat Shock Protein70 Gene and Their Association with Essential Hypertension. Ann Clin Exp Hypertension 4(2): 1041.
INTRODUCTION
Essential hypertension is a global health problem, the prevalence of which continues to increase risk for other cardiovascular diseases. Developing countries particularly India, face a major problem due to the high prevalence of essential hypertension [1]. It is also generally accepted that stress contributes to human high blood pressure. Heat Shock proteins (HSPs) are highly conserved proteins that are expressed in both physiologic and pathologic conditions. HSPs are classified on the basis of molecular weight and the 70 kilo- Dalton proteins referred as the HSP70 family include stress inducible HSP72, HSP70/2 and HSP70/hom [2-4].
Hypertension is a disease with a genetic component and the genetic basis of environmental susceptibility to hypertension may also involve an abnormal control of heat shock protein genes. Previous studies have reported that environmental factors such as diet, stress levels and lifestyle play an important role in genetic predisposition to essential hypertension in humans [5,6]. The genes encoding HSP-70 proteins are mapped within Major histocompatibility complex (MHC) class III region located on chromosome 6. The locus for HSP70s genes, such as HSP 70/1 and HSP 70/2, encoding two identical proteins and the third HSP70/hom gene, are present within the category of human leukocyte antigen (HLA) class III region [7,8]. A Single Nucleotide Polymorphism (SNP) has been identified within the exon region of the HSP70/2 gene at position +1267 (G to A; HSPA1B, OMIM: 603012) [3] and HSP70/1 gene at position +1538 (G to A; HSPA1A, OMIM: 140550) [4]. Another polymorphism has also been described within the codon region of the HSP70/hom gene at position +2437 (C to T; HSPA1L, OMIM: 140559) [4]. The HSP70/1 and HSP70/2 encode an identical heat inducible protein HSP70, whereas HSP70/hom encodes a non-heat inducible form [9]. The HSP70/1 +190G/C polymorphism, a silent substitution which lies 26 base pair (bp) upstream of the translation start site in the 5’ Untranslated region (UTR) region on chromosome 6. The HSP70/2 +1267A/G polymorphism of HSP70-2 lies in the coding region of the gene. The association between variable HSP70/2 messenger RNA (mRNA) expression and the +1267 polymorphism has been reported in diseases, such as insulindependent diabetes mellitus and sepsis [10,11].
Studies have revealed that over-expression of Heat Shock Proteins have cardio protective role and genetic variants of HSP70 would decrease its ability to protect the cells against cardiac ischemia [12-14]. Heat Shock Protein70 polymorphisms have been implicated in various cardiovascular diseases including cardiomyopathy [15], stroke [16] and coronary artery disease (CAD) [17]. The functional polymorphism in HSP70s, such as HSPA1B and HSPA1L was significantly associated with Type 2 diabetes [18]. Increased serum levels of HSP70 are associated with the pathogenesis and chronicity of the diabetes [19]. Most recent study regarding polymorphism in HSPA1B (rs1061581), HSPA1L (rs2227956) and HSPA1 (rs1043618) suggested the gene-disease association of heat shock proteins with lower risk of idiopathic pulmonary fibrosis in Mexican population [20]. Functional SNP in HSPA1A gene (HSP-70/1) is also associated with an increase in risk of coronary heart disease in Han Chinese subjects [21]. Recent study reported the genetic interaction between the polymorphisms of three different genes of HSP70 family (HSPA1A, HSPA1B, and HSPA1L) with increasing risk of essential hypertension in Chinese population [22].
There is no report suggesting the gene-disease association of HSP-70 with essential hypertension in Indian subjects. However, the reported associations have often been inconclusive in other populations. To address this possibility, we investigated the potential involvement of gene variants of HSP70/1, HSP70/2 and HSP70/hom in essential hypertension in Northern Indian population.
MATERIALS AND METHODS
Study population
Sample size (280 patients and 261 controls) is adequate for the present study determined by using standard statistical method in case control group, at the significance level of 0.05 at power 80% on the basis of prevalence of minor alleles referred from previous studies [22]. All the hypertensive patients (males and females) and controls (males and females) were screened randomly from the outpatient department (OPD) clinics of hypertension, Department of Cardiology, All India Institute of Medical Sciences (AIIMS), New Delhi, India. A total of 280 essential hypertensive patients and 261 healthy controls were recruited for a period of three years (2012-2015) for the study, from similar socioeconomic geographical background residing in North India from past three generations. Blood pressure measurements were taken after the patient had been seated on a chair with their feet on the floor and their arms supported at heart level for 10 min. According to Joint National Committee on Prevention, Detection, Evaluation, and Treatment of High Blood Pressure (JNC-VII) the definition of essential hypertension (140 ≤ SBP ≤ 179 mmHg, 90 ≤ DBP ≤ 109 mmHg ) and healthy controls (SBP < 130 mmHg and DBP < 85 mmHg). All hypertensive patients were diagnosed as having essential hypertension and had never been treated with any hypertensive drugs. All the hypertensive patients were not suffering from any known disease, secondary hypertension, diabetes and kidney disease. All procedures performed in present study were in accordance with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. Informed consent was obtained from all participants for being included in the study. Approvals of ethics committee of All India Institute of Medical Sciences (IEC/NP-384/2012), New Delhi and Dr. B. R. Ambedkar Center for Biomedical Research, ACBR, Delhi (F.50-2/ Eth.Com/ACBR/169) were obtained.
Sample collection and processing
Fasting (12 hours) venous blood samples (5 ml) were obtained from all subjects in an ethylene diamine tetra acetic acid (EDTA) vial. Plasma was separated and genomic DNA was extracted from whole blood using the Flexigene DNA kit (QIAGEN®) according to the manufacturer’s instructions.
Genotyping
Genotyping of the three studied polymorphism (HSP70/1:+190G/C (rs1043618), HSP70/2:+1267A/G (rs1061581), and HSP7/hom: +2437T/C (rs2227956),) were identified by using PCR - RFLP techniques. The primers were designed by using GENERUNNER version 3.05 software (Hastings Software Inc. Hastings, NY, USA) based on gene bank sequence (NCBI Reference Sequence: NG_011855.1). PCR amplification conditions were standardized for the above mentioned polymorphisms. The reliability of our genotyping was confirmed by direct sequencing of amplified DNA from randomly selected samples (20%) each from patient and control groups, with no difference observed in results between the two methods.
For HSP70/1 gene polymorphism, the sequence of the primers used were sense primer 5’-ACCTTGCCGTGTTGGAACA-3’ and that of antisense primer was 5’- GGTCTCCGTGACGACTTATA-3’. The amplified PCR product of 337bp was digested with Bsr BI restriction enzyme. The digestion product was separated on 2.5% agarose gel electrophoresis. Digestion of the PCR products yielded bands of 337bp in homozygous wild type (GG), 232bp and 105bp in homozygous rare (CC), and the three bands in the heterozygous (GC) genotypes.
The primers for HSP70/2 gene polymorphism were 5’-ATCTGCGTCTGCTTGGTG-3’ (forward) and 5’- ATCAACGACGGAGACAAG-3’ (reverse). The amplified 998bp products were digested with restriction enzyme Pst I and the digested PCR products were separated on 1.5% agarose gel electrophoresis. Various fragments obtained were 998 bp in homozygous wild type (AA), 770bp and 228bp in homozygous rare (GG) and 998bp, 770bp and 228bp in case of heterozygous rare (AG). Similarly, for HSP70/hom gene polymorphism, the primers used were 5’- CAAGTCTGAGAAGGTACAGGA-3’ (as forward primer) and 5’- GGTAACTTAGATTCAGGTCTG-3’ (as reverse primer). The amplified 876bp products were digested with restriction enzyme NCO I and the digested PCR products were separated on 1.5% agarose gel electrophoresis. Different fragments obtained were 876bp in homozygous wild type (TT), 555bp and 321bp in homozygous rare (CC) and 876bp, 555bp and 321bp in case of heterozygous rare (TC).
Statistical analysis
The sample size was calculated using PS-Power and sample size calculation version 3.0 software, with an α error of 5% and a power 80% and was found to be adequate in the study subjects. Statistical analyses were performed with commercial software (SPSS, ver.11.5; SPSS Inc., Chicago, IL). Normally distributed data are presented as mean± SD. Chi-square goodness of fit was used to verify the agreement of observed genotype frequencies with those expected (Hardy-Weinberg equilibrium). The analysis of variance (ANOVA) was used to calculate the difference between genotype groups using Bonferroni’s method for multiple comparisons between genotype classes. An odds ratio at 95% confidence intervals (CI) was calculated as an index of the association of the genes with the disease. Haplotype analysis and linkage disequilibrium were carried out by Haploview 4.2 to investigate the association between the three polymorphisms of Heat Shock Protein70 family.
RESULTS AND DISCUSSION
Results
Baseline characteristics of the study subjects: The baseline characteristics of patients with essential hypertension (N = 280) and healthy controls (N = 261) are listed in Table (1); Baseline parameters of study subjects. No statistically significant differences in the clinical characteristics were observed between patients and controls. There was non-significant difference between number of male and females in the study subjects. Systolic blood pressures (SBP) in patients were significantly higher (173.69 ± 19.13 mm Hg) than that of controls (115.17 ± 4.25 mm Hg) (p< 0.0001). Similarly, diastolic blood pressures (DBP) in patients were higher (91.09 ± 12.38 mm Hg) than controls (76.25 ± 4.23 mm Hg) (p< 0.0001). High density lipoprotein cholesterol (HDL) and Low density lipoprotein cholesterol (LDL) were comparable in patients and controls.
| Parameters | Patients (280) mean (95%CI)a | Controls (261) mean (95%CI)a | P value* |
| Age (years) | 49.1 ± 12 | 50.2 ± 6 | 0.25 |
| Sex | 168/112 | 167/94 | 0.37 |
| Systolic blood pressure | 173.69 ± 19.13 | 115.17 ± 4.25 | < 0.0001 |
| Diastolic blood pressure | 91.09 ± 12.38 | 76.25 ± 4.23 | < 0.0001 |
| Body mass index(Kg/m2 ) | 25.15 ± 3.54 | 24.80 ± 2.35 | 0.81 |
| Heart Rate (Beat/min) | 73.2 ± 8.7 | 70.2 ± 6.2 | 0.35 |
| Blood Glucose (mg/dl) | 91.8 ± 17.6 | 87.1 ± 15.4 | 0.88 |
| Blood Urea(mg/dl) | 23.6 ± 4.8 | 22.1 ± 4.2 | 0.17 |
| Serum Creatinine | 1.21 ± 0.3 | 0.93 ± 0.22 | 0.21 |
| LDL cholesterol (mg/dl) | 85.8 ± 23.4 | 87.9 ± 29.4 | 0.26 |
| HDL cholesterol (mg/dl) | 44.9 ± 7.2 | 43.1 ± 7.8 | 0.14 |
| Total cholesterol (mg/dl) | 155.8 ± 41.5 | 160.9 ± 37.5 | 0.58 |
| Triglyceride (mg/dl) | 151.0 ± 44 | 145.0 ± 35 | 0.37 |
| Data are means ± SD, Means values of the different parameters in the study subjects were analyzed by chi-square and Student's t-test of significance as appropriate, a 95% confidence interval, Abbreviations: HDL: High-density lipoprotein; LDL: Low-density lipoprotein, * Patients Vs Controls. P < 0.05 is considered to be significant. | |||
Distribution of genotype and allele frequencies of HSP70/ hom, HSP70/1, HSP70/2: Three bi-allelic Single Nucleotide Polymorphisms were investigated: HSP70/1(rs1043618), HSP70/2 (rs1061581) and HSP70/hom (rs2227956) and their genomic locations and related mapping data were obtained from the National Center for Biotechnology Information (NCBI). The genotypic frequencies, allelic frequencies, odds ratio and relative risk estimations for the three polymorphic sites are given in Table (2); Genotype distribution and allele frequency of HSP70 gene
| Polymorphism | Genotype | Hypertensive Patients | Healthy Controls |
| HSP70/hom (rs2227956) | TT | 111 (39.64%) | 134 (51.34%) |
| TC | 143 (51.07%) | 116 (44.44%) | |
| CC | 26 (9.29%) | 11(4.21%) | |
| T allele and C allele | 0.65 & 0.35 | 0.74 & 0.26 | |
| HSP70/1 (rs1043618) | GG | 95 (33.93%) | 115 (44.06%) |
| GC | 147 (52.50%) | 117 (44.83%) | |
| CC | 38 (13.57%) | 29 (11.11%) | |
| G allele and C allele | 0.60 & 0.40 | 0.66 & 0.34 | |
| HSP70/2 (rs1061581) | AA | 111 (39.64%) | 106 (40.61%) |
| AG | 132 (47.14%) | 129 (49.43%) | |
| GG | 37(13.21%) | 26 (9.96%) | |
| A allele and G allele | 0.63 & 0.37 | 0.65 & 0.35 | |
| Genotype data are expressed as number and percentage and allele data as number | |||
polymorphism and Table (3); Association of HSP70 gene polymorphisms with essential hypertension in the study subjects. HSP70/hom showed deviation from the HardyWeinberg equilibrium (HWE) whereas other polymorphisms were found to be in Hardy–Weinberg equilibrium. The genotypes of patient group for HSP70/1 (χ2 = 2.54, p = 0.11), HSP70/2 (χ2 = 0.05, p = 0.82) and HSP70/hom (χ2 = 4.3, p = 0.036) polymorphisms were compared with the genotypes of healthy controls (χ2 = 0.008, p = 0.92; χ2 = 2.16, p = 0.14 and χ2 = 5.31, p = 0.02), respectively. The ‘CC’ genotype of +190G/C/ HSP70/1 was found to be significantly higher in patients (13.57%) with essential hypertension as compared to healthy controls (11.1%). The ‘GG’ genotype of HSP70/2 +1267A/G was found to be 13.21% in patients with essential hypertension whereas 9.96% in healthy controls (Table 2; Genotype distribution and allele frequency of HSP70 gene polymorphism). A statistically significant differences in genotype distribution was found in HSP70/1 +190G/C and HSP70/hom +2437T/C polymorphisms in patients with essential hypertension in comparison to control group with the odds ratio being 1.53 (1.08 – 2.17) & 1.6 (1.14 – 2.25) (Table 3; Association of HSP70 gene polymorphisms with essential hypertension in the study subjects) respectively.
| Polymorphism | Genotype | Chi2 value | Odds ratio | Relative risk | CI (95%) | P value |
| HSP 70/hom | TT Vs TC | 4.94 | 1.48 | 1.22 | 1.04 – 2.11 | 0.020 |
| TT Vs CC | 8.02 | 2.85 | 1.84 | 1.35 – 6.03 | 0.004 | |
| TT Vs TC + CC | 7.46 | 1.60 | 1.27 | 1.14 – 2.25 | 0.006 | |
| T allele Vs. C allele | 8.79 | 1.48 | 1.23 | 1.14 – 1.92 | 0.003 | |
| HSP70/1 | GG Vs GC | 5.1 | 1.52 | 1.23 | 1.05 – 2.18 | 0.023 |
| GG Vs CC | 2.68 | 1.58 | 1.26 | 0.91 – 2.76 | 0.10 | |
| GG Vs GC + CC | 5.84 | 1.53 | 1.24 | 1.08 – 2.17 | 0.015 | |
| G allele Vs. C allele | 4.6 | 1.31 | 1.15 | 1.02 – 1.68 | 0.03 | |
| HSP70/2 | AA Vs AG | 0.02 | 0.97 | 0.98 | 0.68 – 1.40 | 0.88 |
| AA Vs GG | 0.49 | 1.21 | 1.13 | 0.70 – 2.12 | 0.48 | |
| AA Vs AG + GG | 0.01 | 1.02 | 1.0 | 0.72 – 1.44 | 0.92 | |
| A allele Vs. G allele | 0.52 | 1.09 | 1.04 | 0.85 – 1.40 | 0.47 | |
| Patients groups were compared with controls with chi-square (χ2 ) test at one degree of freedom with Odds ratio adjusted for age and sex in both genotypes and alleles. p < 0.05 is considered to be significant. Abbreviations: CI: Confidence Intervals | ||||||
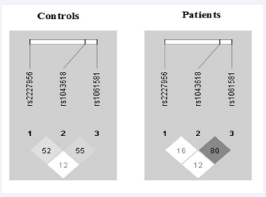
Linkage disequilibrium and haplotype analysis: Linkage disequilibrium and haplotype analysis were carried out for the 3 genotypes of HSP70 using Haploview 4.2 software, to examine for any possible association with the disease phenotype. A significant r2 value of 0.8 was observed for HSP70/1 (rs1043618) and HSP70/2 (rs1061581) genotypes in patients with essential hypertension and in controls r2 value was 0.55 revealing an association between the two polymorphic sites, whereas HSP70/Hom (rs2227956) stands independently and do not shows any association with other polymorphisms (Figure 1); Linkage Disequilibrium of HSP70 gene polymorphism – HSP-70/ HOM, HSP-70/1 & HSP-70/2 in study subjects). The haplotype combination of T-C-G and C-G-A were observed to be higher in patient group (0.216 & 0.208) compared to the controls (0.114 & 0.09) respectively with significant difference (p < 0.001) (Table 4; Haplotype analysis in the study subjects).
| Haplotype | |||||||
| S.N. | HSP70/hom | HSP70/1 | HSP70/2 | Patient frequencies | S.E. | Control frequencies | S.E. |
| H1 | T | G | A | 0.372 | 0.011 | 0.465 | 0.011 |
| H2 | T | G | G | 0.037 | 0.005 | 0.108 | 0.006 |
| H3 | T | C | A | 0.029 | 0.003 | 0.056 | 0.005 |
| H4 | T | C | G | 0.216 | 0.009 | 0.114 | 0.008 |
| H5 | C | G | A | 0.208 | 0.012 | 0.090 | 0.009 |
| H6 | C | G | G | 0.006 | 0.005 | 0.001 | 0.002 |
| H7 | C | C | A | 0.025 | 0.006 | 0.042 | 0.006 |
| H8 | C | C | G | 0.106 | 0.010 | 0.122 | 0.009 |
| The alleles in haplotype are in the order of HSP7/hom:+2437T/C, HSP70/1:+190G/C and HSP70/2:+1267A/G polymorphisms. Abbreviations: SE: Standard Error; SN: Serial Number | |||||||
Figure 1 Linkage Disequilibrium of HSP70 gene polymorphism – HSP70/HOM, HSP-70/1 & HSP-70/2 in study subjects. Figure 1depicts the Linkage Disequilibrium (LD) map, which explains whether the selected SNPs for HSP70 gene polymorphism (HSP-70/HOM, HSP70/1 and HSP70/2) are in LD or inherited together in study subjects. The number shown in figures represents the percentage of correlation between two selected Single Nucleotide Polymorphism (SNPs) for the LD analysis. Linkage disequilibrium plot of SNPs in HSP70/hom (rs2227956), HSP70/1 (rs1043618) and HSP70/2(rs1061581), estimated as ‘r2’ using Haploview 4.2. The plot was built with data of the patients and control group
DISCUSSION
HSP70 is a major stress protein whose production is increased under stress and may be associated with altered protein folding or chaperoning [23]. Several reports suggest an important role of HSP70 proteins in cardiovascular diseases such as hypertrophic cardiomyopathy [15] and ischaemia [24], hypertension [1, 25,26], atherosclerosis [27] and ischemic stroke [28]. Studies also suggest its role in diabetes [18,19] Parkinson’s disease [23,29], high altitude illness [30] and aging [31]. The outcome of various clinical studies support the fact that induction of HSP70 expression in the arterial wall takes place as a physiological response to acute hypertension, such as biomechanical or hemodynamic stress [32-34]. Recent studies suggested that HSP40 acts as an important cofactor for the functional regulation of HSP70s [35], and both the HSPs together play an indispensable role in the chaperone-assisted proteosomal degradation of misfolded proteins [36,37]. The altered expression of HSP70s depends on the sequence variation in the HSP70 gene. It is believed that it may further lead to altered stress tolerance mechanisms and associated pathological states [7]. It has been suggested that the inter-individual differences in HSP70 expression may be related to regulatory mechanisms distinct from transcriptional regulation [11]. Over-expression of HSP70 protects the heart against the damaging effects of ischemia by creatine kinase release, recovery of high energy phosphate stores and correction of metabolic acidosis. The protective effects of HSP70 are ascribed to enhanced protein folding, degradation of abnormal protein, inhibition of apoptosis, protection of cytoskeleton, enhanced NO synthesis [7].
Haplotype analysis is a more informative approach to study the genetic influence on diseases rather than testing isolated genetic markers. In the current study, we used haplotype analysis to explore the genetic contribution of polymorphisms in three genes of HSP70 family to essential hypertension risk in North Indian population. Polymorphism (HSP70/1: +190G/C (rs1043618), HSP70/2:+1267A/G (rs1061581), and HSP70/hom:+2437T/C (rs2227956),) were identified by PCR-RFLP technique. In this study, we observed that the frequency of two haplotypes H4 (T-C-G) and H5 (C-G-A) was higher in patients as compared to the control group. Based on linkage disequilibrium analysis, the combination of HSP70/1 and HSP70/2 polymorphisms that haplotypes (H4 and H5) with two mutant (HSP70/1:+190C and HSP70/2:+1267G) and two wild-type (HSP70/1:+190G and HSP70/2:+1267A) alleles were participated in the increased risk of essential hypertension. The possible combination of haplotypes (H4 and H5) affirmed the existence of interaction between the polymorphisms of HSP70 genes with essential hypertension and may be a factor for the predisposition of disease. Recent linkage study regarding the SNPs in HSP70 gene and essential hypertension also suggested the interaction between haplotypes and their role in increased risk of essential hypertension in Uygur ethnicity of Chinese population [22].
The interaction between genes and environmental factors affecting blood pressure regulation is not completely understood. The presence of these environmental factors on the background of genetic susceptibility might be the determinants of the initiation and progression of hypertension. Since, the environmental factors are generally modifiable; hence their interaction with genetic predisposition to hypertension is of importance to health of an individual. We hypothesize that variations in HSP70 genes/ HSP70 polymorphisms affect the expression of the HSP proteins which may ultimately affect the pathogenesis of essential hypertension. Li and co-workers suggested that circulating HSP70 levels can serve as a biomarker to detect the progression of heart failure [38]. In a 4 year follow up study, high serum levels of HSP70 have been suspected in atheroprotection [39]. Strong proliferative response in peripheral blood lymphocytes of patients with essential hypertension was observed, when HSP70 peptide was used to induce immune tolerance, pointing towards the role of HSP70 in autoimmunity during hypertension and as a key antigen produced by kidney [40]. A previous study has proposed that there is a correlation between serum HSP70 levels and newly diagnosed hypertensive patients [19]. The involvement of HSP70 in cardiovascular diseases, such as essential hypertension and atherosclerosis has been reported earlier [39, 41,42]. Elevated levels of HSP70 are one of the major factors for the predisposition of cardiovascular diseases in established hypertension [25] and transient pregnancy [26]. Strong genetic association between HSP70 gene family and the risk of essential hypertension is well documented [22].
Applying multivariate logistic regression analysis to the baseline parameters documented in Table (1); Baseline parameters of study subjects of the current study, none of the parameters emerged as independent risk factors (data not shown). With respect to HSP70/1 polymorphism, patients with GG genotype were statistically significantly different as compared to healthy individuals. The odds ratio was statistically significant with the risk allele being G, suggesting that ‘G’ allele and its encoded product may play a role in essential hypertension. In addition frequency of G allele was also significantly higher in patients as compared to controls.
In another variant of HSP gene, namely HSP70/hom, we observed a strong association of HSP70/hom polymorphism with the CC genotype and C allele with essential hypertension. The HSP70/hom polymorphism (rs2227956) is associated with the change in amino acid at position 493 from Met to Thr. The amino acid 493 is located in the 18 kDa peptide-binding domain of the beta sheet which constructs the floor of the peptide binding groove [11]. Previous study reported association of increase in frequency of T allele in aged subjects of Irish population [43,44]. The key finding of our study is that HSP-70/hom and HSP70/1 polymorphism is associated with the predisposition to essential hypertension in Northern Indian subjects. The other polymorphism of HSP70/2 was not significantly associated with the essential hypertension in Northern Indian subjects, however these two HSP70 genes (HSP70/1 and HSP70/2) share a similar heat shock factor and also share the same transcriptional start site [9]. It is possible that the HSP70/2 polymorphism is not involved in contributing to essential hypertension, but is in linkage disequilibrium with HSP70/1 gene on chromosome 6. Altered gene or protein expression could be the causal factor for the susceptibility to essential hypertension and the sequence variation might be the attributable reason for the same. To our knowledge, ours is the first study showing a significant association between Heat Shock Protein70/hom, Heat Shock Protein70/1 gene polymorphisms and essential hypertension in North Indian population.
Our study has certain limitations. We only genotyped three polymorphisms in three genes of HSP70 family and did not examine other genes/variants, which might also be associated with hypertension. Other risk factors or intermediate phenotypes such as lifestyles (salt consumption and physical activity) were unavailable for analysis. Although, statistical power of the study was adequate, our results should be considered preliminary and confirmation in a larger sample size with follow-up study is warranted.
CONCLUSION
Taken together, our results supported genetic interaction of the three studied HSP70 polymorphisms with the risk of essential hypertension in North Indian ethnicity. Functional studies are warranted to confirm these findings. To the best of our knowledge, this is the first study to evaluate the polymorphism of HSP70 family of genes in North Indian subjects.
ACKNOWLEDGMENTS
This work was supported by the financial grant [SR/WOS–A/ LS-29/2012(G)] from Department of Science and Technology, India to Dr. Kamna Srivastava.