Recent Antimicrobial Developments Targeting Peptidyl-tRNA Hydrolases
- 1. Department of Chemistry, University of Alabama in Huntsville, USA
ABSTRACT
Peptidyl-tRNA hydrolases (Pths) are essential enzymes found in bacteria, archaea, and eukaryotes. Pths cleave the peptide:tRNA ester bond of peptidyl-tRNAs generated from premature termination of protein synthesis and the expression of short ORFs or minigenes. Accumulation of peptidyl-tRNAs is toxic presumably due to impaired translational initiation or slowed protein synthesis caused by specific tRNA starvation. Pth activity is thus vital for cells to deal with the build-up of peptidyl-tRNAs.
Bacterial genomes encode a highly conserved peptidyl-tRNA hydrolase enzyme, Pth1. Amino acid sequence homology is high across all species and active site residues are strictly conserved. Structures of Pth1 have been solved for several species and catalytically important residues identified. Studies with mini- and partial-substrates have contributed to understanding of substrate recognition. However, the mechanism of action and ability to distinguish between aminoacyl- and peptidyl-tRNAs are not fully understood.
Better understanding of this fundamental biological enzyme has contributed to pharmacological development against this promising new antibacterial target. Expediting the time consuming northern blot assay has paved the way for moderate screening efforts and identification of inhibitory natural product extracts. Computational small molecule docking has also provided initial insight. Overall, Pth1 is a promising new antibacterial target and continued characterization will advance antibiotic development.
KEYWORDS
• Peptidyl-tRNA hydrolase
• Antibiotic
• Antibacterial
CITATION
McFeeters RL (2013) Recent Antimicrobial Developments Targeting Peptidyl-tRNA Hydrolases. JSM Biotechnol Bioeng 1(1): 1006.
INTRODUCTION
Pth Activity is essential and Ubiquitous
Peptidyl-tRNAs are generated when ribosomes abort translation prematurely [1-3], which occurs on average 10% of the time [4]. Peptidyl-tRNAs are then released by ribosome recycling factor and elongation factor-G [4,5], are rescued (i.e. by tmRNA [6] or YaeJ [7,8]), or fall-off at a rate depending on the attached tRNA [9]. Build-up of peptidyl-tRNAs results in tRNA starvation and rapid cell death. Accumulation of peptidyl-tRNAs also results from the expression of minigenes or short ORFs [10- 12]. To avoid excessive build-up of polluting peptidyl-tRNA, it is vital for cells to maintain peptidyl-tRNA hydrolase activity.
Pth activity is universal throughout all kingdoms of life. However, unlike the highly conserved Pth enzymes in bacteria, referred to as Pth1 enzymes, multiple Pths (Pth1, Pth2, and Pth-like domains) are found in archaea and eukaryotes. There is no sequence or structural homology between Pth1 and Pth2 enzymes and the mechanism of bacterial Pth1 action has been demonstrated to be completely different from that of Pth2 [13]. In rabbit reticulocytes, peptidyl-AMP and tRNA-CC are the products of the reaction [14] not just tRNA and peptide. Also, the ability to cleave bacterial initiator fmet-tRNAMet was found in reticulocytes, a property absent in bacterial Pth1s [15,16].
Structures of Pth1 and Pth2 enzymes have been solved. Focusing on Pth1, the crystal structure of the 21 kDa E. coli Pth1 monomer was solved to 1.2 Å [17]. More recently, crystal and solution structures for M. tuberculosis Pth1 have been reported [18,19] along with Pth1 from other pathogenic species including P. aeruginosa [20], F. tularensis [21], M. smegmatis [22], and A. baumannii [23]. As predicted from the high degree of amino acid sequence similarity, all have the same backbone fold. Pth1 family members are globular, single domain proteins that have a central mixed β-sheet surrounded by α-helices (Figure 1).
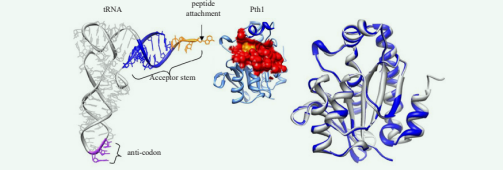
Figure 1 Bacterial Peptidyl-tRNA Hydrolases. Left: tRNAPhe (1EHZ) is shown next to P. aeruginosa Pth1 (4FYJ). The substrate binding site of Pth1 is shown as a red surface with catalytically important His20 in yellow. Helix-4 is in dark blue. The unpaired region of the tRNA acceptor stem is shown in orange. Right: Ribbon diagram of Pth1 from P. aeruginosa (blue, 4FYJ) aligned with E. coli (white, 2PTH). Heavy backbone atom RMSD is 0.63 Å.
Helix-4 occludes three residues, N10, H20, and D93 (as numbered in E. coli Pth1), identified by site directed mutagenesis to be crucially involved in enzyme activity [24]. 15N NMR relaxation data suggest the putative active site and the helix-4 cover exhibit motions on the millisecond to microsecond timescale, thought to be linked to interaction with the peptidyl-tRNA substrate [18].
Speculation on the enzymatic mechanism include H20 acting as a catalytic base. Residues K105 and R133 are suggested to be involved in clamping the 5’-phosphate group of the tRNA moiety [17,24-26]. Structural information is needed to resolve these postulates, yet no structure of the enzyme-substrate complex has been determined for any Pth.
Pth1 is a Promising Target for Antibacterial Development
With the relentless development of drug resistance and re-emergence of many pathogenic bacteria, the need for new antibiotics and new antibiotic targets is urgent and growing [27,28]. It is well known that the number of antimicrobial agents being brought to market has steadily declined [29]. Pth1 provides a much needed avenue for novel antibacterial development. The essential function, high conservation across bacterial species, and lack of essential human equivalent make Pth1 a prime target for antibacterial development. The high amino acid sequence conservation across bacterial species suggests a small molecule inhibitor found against one is likely to work against many others. Thus, broad spectrum inhibitors targeting Pth1 are a distinct possibility.
Targeting a new essential bacterial enzyme, Pth1 inhibitors will be effective against drug resistant strains and provide the potential for fresh, new, and original combinatorial therapies. Although bacteria have but a single class of Pth enzyme, eukaryotes possess several structurally unrelated Pth enzymes that subsume peptidyl-tRNA hydrolase activity. Pth1 knock out does not alter cell viability in yeast [30], thus Pth1 does not appear to be essential in eukaryotes. The very different structure for Pth2 [13,31-33] and Pth-domain containing proteins [34,35] mitigates concern for mitochondrial Pth impairment.
Molecular modeling studies indicate two proximal binding sites for small molecules on either side of the catalytically important H20 residue, in the crevice containing the putative active site [18]. The next step in targeted inhibition is already underway. Natural product screening shows promise for revealing Pth1 specific inhibitors [36,37]. Also, screening of a custom combinatorial library has also revealed a family of compounds that show potent in vitro and in vivo inhibition. Both show that Pth1 specific inhibition is possible, not inhibiting Pth2 in vitro.
CONCLUDING REMARKS
The solid structural biology foundation and first identification of specific small molecule inhibition of Pth1 are beginning to bridge the gap between in vitro biochemical studies and lead compound identification. Such translational progress will continue, aided by medicinal chemistry and computational studies further contributing to small molecule binding studies. Overall, Pth1 is a promising new antibacterial target and comprehensive characterization will significantly advance structural biology of the system, understanding of the Pth:peptidyl-tRNA interaction, and possibly development of next generation antibiotics.
REFERENCES
- Jørgensen F, Kurland CG. Processivity errors of gene expression in Escherichia coli. J Mol Biol. 1990; 215: 511-521.
- Manley JL. Synthesis and degradation of termination and premature-termination fragments of beta-galactosidase in vitro and in vivo. J Mol Biol. 1978; 125: 407-432.
- Kurland CG, Ehrenberg M. Constraints on the accuracy of messenger RNA movement. Q Rev Biophys. 1985; 18: 423-450.
- Heurgué-Hamard V, Karimi R, Mora L, MacDougall J, Leboeuf C, Grentzmann G, et al. Ribosome release factor RF4 and termination factor RF3 are involved in dissociation of peptidyl-tRNA from the ribosome. EMBO J. 1998; 17: 808-816.
- Karimi R, Pavlov MY, Heurgué-Hamard V, Buckingham RH, Ehrenberg M. Initiation factors IF1 and IF2 synergistically remove peptidyl-tRNAs with short polypeptides from the P-site of translating Escherichia coli ribosomes. J Mol Biol. 1998; 281: 241-252.
- Singh NS, Varshney U. A physiological connection between tmRNA and peptidyl-tRNA hydrolase functions in Escherichia coli. Nucleic Acids Res. 2004; 32: 6028-6037.
- Handa Y, Inaho N, Nameki N. YaeJ is a novel ribosome-associated protein in Escherichia coli that can hydrolyze peptidyl-tRNA on stalled ribosomes. Nucleic Acids Res. 2011; 39: 1739-1748.
- Gagnon MG, Seetharaman SV, Bulkley D, Steitz TA. Structural basis for the rescue of stalled ribosomes: structure of YaeJ bound to the ribosome. Science. 2012; 335: 1370-1372.
- Menninger JR. The accumulation as peptidyl-transfer RNA of isoaccepting transfer RNA families in Escherichia coli with temperature-sensitive peptidyl-transfer RNA hydrolase. J Biol Chem. 1978; 253: 6808-6813.
- Cruz-Vera LR, Hernandez-Ramon E, Perez-Zamorano B, Guarneros G. The rate of peptidyl-tRNA dissociation from the ribosome during minigene expression depends on the nature of the last decoding interaction. J Biol Chem. 2003; 278: 26065-26070.
- Hernández-Sánchez J, Valadez JG, Herrera JV, Ontiveros C, Guarneros G. lambda bar minigene-mediated inhibition of protein synthesis involves accumulation of peptidyl-tRNA and starvation for tRNA. EMBO J. 1998; 17: 3758-3765.
- Tenson T, Herrera JV, Kloss P, Guarneros G, Mankin AS. Inhibition of translation and cell growth by minigene expression. J Bacteriol. 1999; 181: 1617-1622.
- Fromant M, Schmitt E, Mechulam Y, Lazennec C, Plateau P, Blanquet S. Crystal structure at 1.8 A resolution and identification of active site residues of Sulfolobus solfataricus peptidyl-tRNA hydrolase. Biochemistry. 2005; 44: 4294-4301.
- Gross M, Crow P, White J. The site of hydrolysis by rabbit reticulocyte peptidyl-tRNA hydrolase is the 3'-AMP terminus of susceptible tRNA substrates. J Biol Chem. 1992; 267: 2080-2086.
- Schulman LH, Pelka H. The structural basis for the resistance of Escherichia coli formylmethionyl transfer ribonucleic acid to cleavage by Escherichia coli peptidyl transfer ribonucleic acid hydrolase. J Biol Chem. 1975; 250: 542-547.
- Dutka S, Meinnel T, Lazennec C, Mechulam Y, Blanquet S. Role of the 1-72 base pair in tRNAs for the activity of Escherichia coli peptidyl-tRNA hydrolase. Nucleic Acids Res. 1993; 21: 4025-4030.
- Schmitt E, Fromant M, Plateau P, Mechulam Y, Blanquet S. Crystallization and preliminary X-ray analysis of Escherichia coli peptidyl-tRNA hydrolase. Proteins. 1997; 28: 135-136.
- Pulavarti SV, Jain A, Pathak PP, Mahmood A, Arora A. Solution structure and dynamics of peptidyl-tRNA hydrolase from Mycobacterium tuberculosis H37Rv. J Mol Biol. 2008; 378: 165-177.
- Selvaraj M, Roy S, Singh NS, Sangeetha R, Varshney U, Vijayan M. Structural plasticity and enzyme action: crystal structures of mycobacterium tuberculosis peptidyl-tRNA hydrolase. J Mol Biol. 2007; 372: 186-193.
- Hughes RC, McFeeters H, Coates L, McFeeters RL. Recombinant production, crystallization and X-ray crystallographic structure determination of the peptidyl-tRNA hydrolase of Pseudomonas aeruginosa. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2012; 68: 1472-1476.
- Clarke TE, Romanov V, Lam R, Gothe SA, Peddi SR, Razumova EB, et al. Structure of Francisella tularensis peptidyl-tRNA hydrolase. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2011; 67: 446-449.
- Kumar A, Singh N, Yadav R, Kumar RP, Sharma S, Arora A, et al. Crystal structure of peptidyl-tRNA hydrolase from mycobacterium smegmatis reveals novel features related to enzyme dynamics. Int J Biochem Mol Biol. 2012; 3: 58-69.
- Kaushik S, Singh N, Yamini S, Singh A, Sinha M, Arora A, et al. The Mode of Inhibitor Binding to Peptidyl-tRNA Hydrolase: Binding Studies and Structure Determination of Unbound and Bound Peptidyl-tRNA Hydrolase from Acinetobacter baumannii. PLoS One. 2013; 8: e67547.
- Goodall JJ, Chen GJ, Page MG. Essential role of histidine 20 in the catalytic mechanism of Escherichia coli peptidyl-tRNA hydrolase. Biochemistry. 2004; 43: 4583-4591.
- Ostheimer GJ, Hadjivassiliou H, Kloer DP, Barkan A, Matthews BW. Structural analysis of the group II intron splicing factor CRS2 yields insights into its protein and RNA interaction surfaces. J Mol Biol. 2005; 345: 51-68.
- Ito K, Murakami R, Mochizuki M, Qi H, Shimizu Y, Miura K, et al. Structural basis for the substrate recognition and catalysis of peptidyl-tRNA hydrolase. Nucleic Acids Res. 2012; 40: 10521-10531.
- Morens DM, Folkers GK, Fauci AS. The challenge of emerging and re-emerging infectious diseases. Nature. 2004; 430: 242-249.
- Morens DM, Folkers GK, Fauci AS. Emerging infections: a perpetual challenge. Lancet Infect Dis. 2008; 8: 710-719.
- Boucher HW, Talbot GH, Bradley JS, Edwards JE, Gilbert D, Rice LB, et al. Bad bugs, no drugs: no ESKAPE! An update from the Infectious Diseases Society of America. Clin Infect Dis. 2009; 48: 1-12.
- Rosas-Sandoval G, Ambrogelly A, Rinehart J, Wei D, Cruz-Vera LR, Graham DE, et al. Orthologs of a novel archaeal and of the bacterial peptidyl-tRNA hydrolase are nonessential in yeast. Proc Natl Acad Sci U S A. 2002; 99: 16707-16712.
- De Pereda JM, Waas WF, Jan Y, Ruoslahti E, Schimmel P, Pascual J. Crystal structure of a human peptidyl-tRNA hydrolase reveals a new fold and suggests basis for a bifunctional activity. J Biol Chem. 2004; 279: 8111-8115.
- Powers R, Mirkovic N, Goldsmith-Fischman S, Acton TB, Chiang Y, Huang YJ, et al. Solution structure of Archaeglobus fulgidis peptidyl-tRNA hydrolase (Pth2) provides evidence for an extensive conserved family of Pth2 enzymes in archea, bacteria, and eukaryotes. Protein Sci. 2005; 14: 2849-61.
- Shimizu K, Kuroishi C, Sugahara M, Kunishima N. Structure of peptidyl-tRNA hydrolase 2 from Pyrococcus horikoshii OT3: insight into the functional role of its dimeric state. Acta Crystallogr D Biol Crystallogr. 2008; 64: 444-453.
- Singarapu KK, Xiao R, Acton T, Rost B, Montelione GT, Szyperski T. NMR structure of the peptidyl-tRNA hydrolase domain from Pseudomonas syringae expands the structural coverage of the hydrolysis domains of class 1 peptide chain release factors. Proteins. 2008; 71: 1027-1031.
- Handa Y, Hikawa Y, Tochio N, Kogure H, Inoue M, Koshiba S, et al. Solution structure of the catalytic domain of the mitochondrial protein ICT1 that is essential for cell vitality. J Mol Biol. 2010; 404: 260-273.
- Harris SM, McFeeters H, Ogungbe IV, Cruz-Vera LR, Setzer WN, Jackes BR, et al. Peptidyl-tRNA hydrolase screening combined with molecular docking reveals the antibiotic potential of Syzygium johnsonii bark extract. Nat Prod Commun. 2011; 6: 1421-1424.
- McFeeters H, Gilbert MJ, Thompson RM, Setzer WN, Cruz-Vera LR, McFeeters RL. Inhibition of essential bacterial peptidyl-tRNA hydrolase activity by tropical plant extracts. Nat Prod Commun. 2012; 7: 1107-1110.






































































































































