Bacterial Genomic DNA Mediated Phagocytosis of Apoptotic Neutrophils by Macrophages without Provoking Inflammation
- 1. Department of geriatrics Pulmonary, Anhui Medical University, China
Abstract
The bacterial genomic DNA from antibiotics killed bacteria may have unique functions in the pathogenesis of bacterial infectious diseases. We investigate the ability of macrophages to engulf the apoptotic neutrophils mediated by bacterial genomic DNA (B-DNA). We found that the percentage of macrophages with ingested apoptotic neutrophils was increased after 2µg/ml B-DNA stimulation. The enhancement was consistent with cell surface and total TLR9 expression on macrophages. B-DNA had no significant effect on the secretion of IL-6 and TNF-α from macrophages after B-DNA stimulation and ingestion of apoptotic neutrophils. We proposed that bacterial genomic DNA released from the antibiotic killed bacteria could mediate phagocytosis of apoptotic neutrophils by macrophages through TLR-9 signaling pathway in bacterial infectious diseases without provoking inflammation, which is one important process of the immune system and is necessary for the homeostatic maintenance.
CITATION
Jiong W, Huimin C, Jin Y, Xuexue P, Xueqin J, et al. (2015) Bacterial Genomic DNA Mediated Phagocytosis of Apoptotic Neutrophils by Macrophages without Provoking Inflammation. JSM Cell Dev Biol 3(1): 1016.
Keywords
Bacterial genomic DNA; Macrophage; Apoptosis; Phagocytosis; Inflammation.
INTRODUCTION
bacterial genomic DNA released from antibiotics killed bacteria in vivo and in vitro. However they may have unique functions Respiratory tract bacterial infections, characterized by neutrophil recruitment and accumulation of protein-rich edema fluid resulting impaired lung function, cause a globally significant disease burden and account for a high morbidity and mortality in many undeveloped countries [1,2]. Neutrophils are the most abundant leukocyte population in human blood and are rapidly mobilized to sites of infectious respiratory tract. Fighting and destroying bacterial infections are the primary functions of neutrophils [3,4]. Once the neutrophils have completed their functions, an orderly elimination is essential for the resolution of inflammation [5,6]. The latter process occurs by the apoptosis of neutrophils in situ followed by their rapid engulfment and degradation by macrophages and other phagocytes [7,8]. Any kind of impaired or defective clearance of apoptotic neutrophils during inflammation would lead to continuous or repeated bouts of acute inflammation, resulting in ongoing tissue damages [9,10].
When bacteria are killed by antibiotics, the bacterial genomic DNA were continuously released through decomposition of organic matter in vivo and then degraded by human endogenous DNAases. So far little focus has been on what the effects of in the pathogenesis of bacterial infectious diseases. We have demonstrated that artificially synthesized unmethylated CpG oligodeoxynucleotides could enhance the ingestion of apoptotic neutrophils by macrophages without increase of pro inflammatory cytokines release [11,12]. In the present study, we evaluated the ability of macrophages to phagocyte apoptotic neutrophils activated by bacterial genomic DNA in vitro.
MATERIALS AND METHODS
Materials
Percoll was purchased from Amersham Biosciences and Annexin V-FITC Apoptosis Detection kit was from BD Bioscience, Dulbecco’s modified Eagle’s medium (DMEM), fetal bovine serum were obtained from Invitrogen; IL-6 and TNF-α radio immune assay (RIA) kits were from the General Hospital of PLA Institute of Science and Technology Developmental Center. O-dianisidine dihydrochloride, 3-(4,5-dimethylthiozol-2-yl) 2,5-diphenyltetrazolium bromide (MTT) was from Sigma; Anti-mouse TLR-9 polyclonal antibody was purchased from eBioscience. RAW264.7 cells and Escherichia Coli JM109 strains were preserved by our own laboratory.
MAIN METHODS
Preparation of bacterial DNA
Cultured Escherichia Coli JM109 in Luria-bertani (LB) Broth at 37°C overnight was pelleted by centrifugation at 10,000×g for 1 minute. The pelletes were lysed in buffer consisting of 50 mM Tris-HCl-5 mM EDTA (pH 8.0); proteinase K (0.2 mg/ml, PH7.5) and sodium dodecyl sulfate (0.5%) and incubated at 50°C overnight. DNA was purified from the lysate by repeated extraction with phenol-chloroform-isoamyl alcohol, precipitated with sodium acetate and ethanol, and then was dissolved in water and stored at -20°C in aliquots [13,14]. The concentration of DNA was measured in a photometer at 260nm.
Assay of cytoxicity of B-DNA on RAW264.7 cells
The cytoxicity of extracted bacterial genomic DNA on in vitro cultured RAW264.7 cells was evaluated by using the MTT colorimetric assay. Briefly, RAW264.7 cells were plated in 96 wells and incubated overnight. After 24 hours stimulation of 2µg/ml B-DNA supplemented with or without chloroquine for 2h ahead in triplicate, cells were washed twice in PBS, 180 μl of fresh medium and 20 μl of MTT (5 mg/ml) were added to each well, and the plate was incubated for an additional 4 h at 37°. The medium was removed, and the MTT crystals were completely solubilized with libration for 10 min with 50% DMSO. Spectrophotometric absorbance of each sample was then measured at wavelengths A570 nm. The percentage of cell viability was calculated using the following equation: viability (%) = OD treated group ×100/ODcontrol group .
Isolation and induction of apoptosis of neutrophils
10ml heparinized peripheral venous blood was obtained from healthy donor with written informed consent and approved by the medical ethic committee of the first affiliated hospital of Anhui Medical University. Neutrophils were isolated using the Percoll discontinuous density gradient centrifugation as previously described with some modification [15,16]. Briefly, 10ml heparinized peripheral venous blood was incubated for 5 min in 50 ml red blood cell lysis buffer (170 mM NH4 Cl, 10 mM KHCO3 , 0.1 mM EDTA, pH 7.3) to remove red blood cells. The remaining cells were then layered on the top of Percoll discontinuous density gradient and centrifuged (1000g, 30min, room temperature) without braking. Then isolated neutrophils were washed three times, cultured at a density of 1 x 106 cells/ ml in DMEM supplemented with 10% fetal calf serum (FCS), 2 mmol/L glutamine, 100 IU/ml penicillin and 100 mg/ml streptomycin. The cell viability and purity of neutrophils isolated using this method were ≥95% by Wright’s staining and trypan blue staining respectively. Spontaneous apoptosis was achieved by culturing for 20h [17,18]. The percentage of apoptotic neutrophils assessed by Annexin V-FITC apoptosis detection kit by flow cytometry was 25 60% while necrosis was less than 3%.
Phagocytosis assessment
For phagocytosis, 2× 104 RAW264.7 cells were transferred into 24-well plates in each well in 500µl DMEM with 10% FCS and allowed to grow overnight. After another 24 hours treatment with 2µg/ml B-DNA supplemented with or without chloroquine for 2h ahead, the cells were randomly divided into 3 groups: normal control group, B-DNA group, Chloroquine+B DNA group and then co-incubated with apoptotic neutrophils (4×104 /well) at 37°C for 1h. The wells were washed vigorously with phosphate-buffered saline (PBS) with 0.02mol/L EDTA to remove non-ingested neutrophils. RAW264.7 cells were fixed in 2.5% glutaraldehyde in PBS and stained with haematoxylin and the ingested neutrophils were stained by o-dianisidine for light microscope [19,20]. All experiments were performed in duplicate and at least 200 cells were counted in randomly selected fields.
Measurement of Total TLR9 expression
For measuring the total TLR9 expression (including intracellular and cell surface), the cells were transferred to a 35mm dish at a density of 2× 106cells/dish in 1000μL complete medium and cultured for 24 hours. Macrophages were activated by 2µg/ml B-DNA for 24h supplemented with or without chloroquine for 2h ahead. Then the cells were harvested and lysed on ice for 10min in lysis buffer. Cell lysates were then subjected to Western blotting for TLR9 as previously described [21,22]. Briefly, Proteins were separated by SDS-PAGE and electro transferred to a polyvinylidene fluoride membrane, followed by blocking with 5% nonfat dry milk in PBS. After washing with PBS containing 0.1% Tween 20, the membrane was incubated with rabbit anti-mouse TLR-9 antibody (1:250 dilutions) in 1% milk for 1 hour at room temperature. After repeated washing, the membrane was incubated with the appropriate secondary antibody and signal was detected with Pierce ECL plus Western Blotting Detection Reagents. Bands density was measured using an imaging densitometer for quantitation. Equal protein loading was confirmed with α-tubulin antibody.
Measurement of cell surface TLR9 expression of macrophages
For measuring the surface expression of TLR9 on macrophages, the cells were incubated with the appropriately diluted anti TLR9 mAb (eBioscience) after 24 hours stimulation of 2µg/ml B-DNA supplemented with or without 2μg/ml chloroquine for 2h ahead, followed by incubation with Cy3–conjugated secondary mAb (Sigma).Then macrophages were analyzed for TLR9 with an EPICS XL flow cytometer (Beckman Coulter, Fullerton, CA).
Production analysis of IL-6and TNF-α
To analyze the secretion of pro-inflammatory cytokines from macrophages after B-DNA activation and ingestion of apoptotic neutrophils, macrophages were cultured in DMEM in 48-well plates and allowed to grow overnight. After 24 hours stimulation of 2µg/ml B-DNA supplemented with or without 2μg/ml chloroquine for 2h ahead, the supernatants were collected and stored. The cells were then incubated together with apoptotic neutrophils at 37°C. After 60 min, macrophages were carefully washed, and 250μl DMEM was added to each well. The medium was harvested and refreshed after 4, 8, and 12, 24h incubation, centrifuged, and stored at -80°C. The concentrations of IL-6 and TNF-α were measured by radio-immune assay (RIA) according to the manufacturer’s instruction. Measurements were performed in duplicate.
Statistical analysis
All experiments were performed for three or five independent times. Data were expressed as the means ± standard error and were analyzed for significant differences by unpaired Student’s t test and one-way ANOVA with SPSS 10.0. Differences were considered statistically significant if P value <0.05.
RESULTS
Indicated concentration of B-DNA was innoxious to macrophages
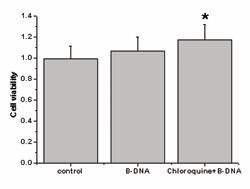
As shown in Figure 1, 2µg/ml B-DNA used in the experiments was innoxious to macrophages evaluated by MTT test.
Figure 1: Indicated concentrations of B-DNA was innoxious to macrophages. Assessment of cell viability by MTT assay 2µg/ ml B-DNA was innoxious, pre-treatment with 2μg/ml B-DNA+CHQ promoted the proliferative capacity of macrophages compared with the control groups (p<0.05). Data were expressed as mean± SEM (n=5).
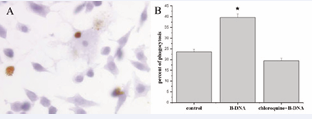
However, pre-treatment with 2μg/ml B-DNA+CHQ promoted the proliferative capacity of macrophages compared with the control groups (p<0.05), which indicated that they also were innoxious to macrophages. B-DNA up-regulated ingestion of apoptotic neutrophils by macrophages As shown in Figure 2A, the engulfed apoptotic neutrophils were visualized as brown staining peroxidase-positive cells inside the blue macrophages which were counter-stained with haematoxylin. Figure 2B showed the uptake of apoptotic neutrophils by macrophages activated by B-DNA.
Figure 2: B-DNA up-regulated ingestion of apoptotic neutrophils. Analysis of up-take of apoptotic neutrophils by macrophages. A: Photomicrograph of macrophages with apoptotic neutrophils. Engulfed apoptotic neutrophils were seen as condensed peroxidase positive brown staining bodies within the blue macrophages which were counter-stained with haematoxylin. B: The effect of B-DNA on the ingestion of apoptotic neutrophils by macrophages. Data were expressed as percentage of phagocytosis± S.E. *p<0.05 vs control group.
The percentages of macrophages with apoptotic neutrophils were 23.6%, 39.58% and 19.40% in control, B-DNA and B-DNA+CHQ groups respectively. B-DNA significantly enhanced uptake of apoptotic neutrophils by macrophages compared with the control groups (p<0.05), however, the effect was blocked by CHQ pretreatment.
Total TLR9 protein was up-regulated after B-DNA treatment
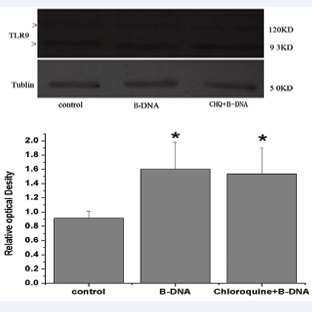
As shown in Figure 3, the total TLR9 expression was stronger after 2µg/ml B-DNA treatment in macrophages while chloroquine had no significant effect on the up-regulated expression of TLR9 induced by B-DNA treatment.
Figure 3: Total TLR9 protein was activated by B-DNA. Analysis of total TLR-9 expression in macrophages by Western blot. A. Two isoforms of the TLR9 protein (93KD and 120KD) were indicated, Western blot of a-tubulin (50KD) was performed to confirm equal loading of protein on the gel. B. Relative optical densitometric values were expressed as mean± SEM (n=3). *p<0.05 vs control group.
B-DNA pretreatment up-regulated surface TLR9
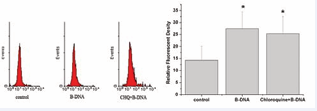
As shown in Figure 4, cell surface TLR9 expression was also up-regulated after B-DNA pre-treatment and chloroquine had no significant effect on the enhanced expression induced by B-DNA activation. The cell surface TLR9 expression was in step with the total TLR9 expression of macrophages.
Figure 4: The cell surface TLR9 expression was also increased after B-DNA pretreatment. Analysis of cell surface TLR9 expression on macrophages using flow cytometer. A. Representative flow cytometry analysis of cell surface TLR9 expression on macrophages. B. Mean fluorescent density data were expressed as mean± SEM (n=5). *p<0.05 vs control group.
B-DNA had no effect on the production of IL-6 and TNF-α
As shown in Table 1, the concentrations of IL-6 and TNF-α protein in the supernatants of cultured macrophages were not affected after B-DNA stimulation and ingestion of apoptotic neutrophils as time lapsed.
|
Table 1: B-DNA had no effect on the secretion of IL-6 and TNF-α after B-DNA activation and ingestion of apoptotic neutrophils. |
||
|
Incubation time |
IL-6(pg/ml) |
TNF-?(ng/ml) |
|
0h 4h 8h 12h 24h |
21.5±0.63 34.7±1.32 40.5±1.44 51.3±0.83 48.2±0.72 |
0.45±0.23 0.35±0.13 0.62±0.16 0.50±0.03 0.65±0.23 |
|
The concentrations of IL-6 and TNF-α in the medieum of macrophages were not affected after B-DNA stimulation and ingestion of apoptotic neutrophils as time lapsed. Data were expressed as mean± SEM (n=5). |
||
DISCUSSION
In respiratory tract bacterial infectious diseases, when bacteria are killed by antibiotics, DNA are released into bloodstream continuously and then degraded by human endogenous DNAases. Alexandra heininger et al [23,24] demonstrated that antibiotic killed bacteria do not release a large proportion of their DNA into the bloodstream and that the released bacteria DNA is not readily degraded by blood DNAases. Little focus has been on what the bacterial DNA acted after they were released before their degradation. In the present study, we investigated the ability of macrophages to ingest apoptotic neutrophils activated by bacterial genomic DNA and the production of pro-inflammatory cytokines after B-DNA stimulation. We found that B-DNA up regulated the ingestion of apoptotic neutrophils by macrophages. Tumor necrosis factor-α (TNF-α) and interleukin-6 (IL-6) are pivotal for provoking inflammation and are considered to be crucial to the later immune response [25,26]. In the study, we found that the production of TNF-α and IL-6 were not affected in the supernatants of macrophages after B-DNA stimulation and ingestion of apoptotic neutrophils. These results suggested that B-DNA enhanced ingestion of apoptotic neutrophils without provoking inflammation which is important for the restriction and resolution of inflammation [27, 28].
Toll-like receptor 9 (TLR9) is an innate immune sensor for microbial DNA that erroneously responds to self DNA in autoimmune disease and TLR9 is normally expressed in endosomes/lysosomes where it is activated by pathogen-derived DNA [29,30].Macrophage has transmembrane receptors that bind B-DNA for engulfment, and then fused with cytomembrane [31,32]. The combination was transported to tubular lysosomal compartment where the endosome formed. Chloroquine can prevent the acidification of endosomes by raising the endosomal PH and interfere with cell cytotoxicity presumably by inhibiting lysosomal enzyme function [33,34]. In the present study, we found that enhanced phagocytic ability of macrophages activated by B-DNA was abolished when the cells were pretreated with CHQ; CHQ had no effect on TLR9 expression. Enhancement of phagocytosis of apoptotic neutrophils by macrophages was in step with cell surface and total TLR9 expression in macrophages. These results suggested that increase of apoptotic neutrophils uptake by macrophages activated by B-DNA also requires endosomal acidification/maturation. These observations were consistent with previous studies [12,35,36]. However more research is needed to understand the effects of B-DNA were mediated through surface or intracellular TLR9 or by which types of TLR9 isoforms.
In conclusion, we proposed that bacterial genomic DNA released from the antibiotic killed bacteria could enhance the ingestion of apoptotic neutrophils by macrophages through TLR-9 signaling pathway in bacterial infectious diseases without provoking inflammation, which is one important process of the immune system and is necessary for the homeostatic maintenance.