Cellular Engineering for Enhanced Production of Potent Antimalarial Drug: Artemisinin
- 1. Department of Green technology, Ipca Laboratories ltd., Sejavta-457002, Dist. Ratlam, Madhya Pradesh, india
- 2. Division of Gastroenterology and Liver Disease, Case Western Reserve University, Cleveland, Ohio, USA
- 3. Departement of Biotechnology, Faculty of Science, Jamia Hamdard, Hamdard Nagar, New Delhi-110062, India
Abstract
Improvement in the yield of artemisinin, a novel and potent anti malarial compound, through conventional breeding, in-vitro culture, hairy root culture, cell suspension culture and total organic synthesis are still a challenge. Genetic engineering tools, however, can be employed to over express genes coding for enzymes associated with the rate limiting steps of artemisinin biosynthesis or to inhibit the enzymes of other pathway competing for its precursors. Based on genetic manipulations of the pathway and cloning of the related genes, more recently cellular engineering of artemisinin biosynthesis has been envisaged in Artemisia annua L., Cichorium intybus L. and microbes aiming at increasing artemisinin content in transgenic organism for the past few years. The strategies which can be employed to enhance the yield of artemisinin both in vivo and in vitro are discussed in the present review.
Keywords
Cellular engineering; Artemisinin; A. annua L. HMG-CoA reductase; Amorpha-4, 11-diene synthase; CYP71AV1; Squalene synthase; Farnesyl diphosphate synthase; E. coli; S. cerevisiae.
Citation
Ram M, Niranjan R, Jain DC, Abdin MZ (2013) Cellular Engineering for Enhanced Production of Potent Antimalarial Drug: Artemisinin. JSM Cell Dev Biol 1(1): 1002.
INTRODUCTION
Malaria is causing more than a million deaths and 500 million clinical cases annually. Despite tremendous efforts for the control of malaria, the global morbidity and mortality have not been significantly changed in the last 50 years [1]. The key problem is the failure to find effective medicines against malaria. Artemisinin, a sesquiterpene lactone containing an endoperoxide bridge obtained from a Chinese medicinal plant Artemisia annua L., has been demonstrated as an effective and safe alternative therapy against malaria [2]. Artemisinin and its derivatives are found effective against multi-drug resistant Plasmodium sp., which is especially prevalent in Southeast Asia, South America and more recently in Africa [3-5]. It is also found effective against other infectious diseases such as schistosomiasis, HIV, hepatitis B and leishmmaniasis [6-10]. It has also been reported to be effective against a variety of cancer cell lines including breast cancer, human leukemia, colon cancer and small cell-lung carcinomas [11-13]. Unfortunately in some parts of CombodiaThailand border, malaria parasite (P. falciparum) has developed resistant against artemisinin monotherapies [14]. To combat this problem WHO now has recommended use of in artemisinin based-combination therapy (ACT). The exponential increase in the number of countries adopting ACTs has led to a rapid increase in demand (180 metric tons year-1) for artemisinin and its derivatives. However, its global production (120 metric tons year-1) that is far behind from its global demand (180 metric tons year-1) [15]. The relatively low yield of artemisinin in A. annua L. leaves (0.01-1.1%) and unavailability of economical, viable biotechnological or synthetic protocol are however, the major limitations to commercialization of drug [16-18] [67]. It is empirical to enhance the production of artemisinin to bring down the cost of ACT treatment and make it affordable to developing Countries. To overcome this 48 problem, efforts are being made worldwide to enhance its production employing various approaches such as conventional breeding, biochemical, physiological, molecular and hairy root culture techniques [19,24,84,91,103,119,120]. These approaches show potential for future development, but improvements delivered by them so far have not met the global demand.
Cellular engineering aims at to (i) modify cellular metabolite composition in order to produce new compounds, (ii) enhance the production of existing compounds, and (iii) eliminate the undesirable ones. Plant metabolism is modified either by introducing novel genes or pathways, or enhancing the expression of endogenous pathways e.g., by up-regulating transcription factors. Further, down-regulation of endogenous genes to suppress or block the production of undesirable metabolites is accomplished by silencing target genes with anti-sense expression or RNA interference (RNAi) [46]. The advancements and limitations of genetic modification of plants have been regularly overviewed in numerous reviews and commentaries [30,35,44,47,86]. In addition, altering metabolic enzymes or pathways has become an important approach for investigating cell physiology [52]. Application of cellular engineering for the production of artemisinin and its precursors, particularly in Artemisia annua L., Cichorium intybus L. and microbes, has been adapted very recently [24,56,78,81,84,87,91,120]. The production of precursors of artemisinin such as amorpha-4, 11-diene and artemisinic acid, especially in E. coli and yeast, has become a prime example of the capabilities of this parts-listand-systems-design approach to microbial genetic engineering. Initially, the production of the FDP precursor was optimized, and then enzyme catalyzing the first committed step in the artemisinin pathway i.e. amorpha-4, 11-diene synthase was overexpressed in E. coli [78]. Several enzymes involved in the early steps of artemisinin biosynthesis have been discovered which include HMGR, FPS, ADS, DXS, DXR, SQS, CYP71AV1 etc. (Table 1).
Table 1: Genes related to artemisinin biosynthesis in A. annua L.
| Enzyme | Gene | Function | Location | Gene Bank accession no. | Reference |
| Deoxyxylulose synthae | dxs | 1-Deoxy-D-xylulose-5-phosphate synthase activity | Plastid | AF182286 | Souret etal., 2002 |
| Deoxyxylulose reductase | Dxr | Isomerase and oxidoreductase activity | Plastid | AF182287 | Souret etal., 2002 |
| 3-Hydroxy-3-methyl | hmg r | Catalyse the two step reduction of | Cytosol | Souret etal., 2002 | |
| glutaryl coenzyme A | S-HMG-CoA into R-mevalonate | ||||
| Farnesyl diphosphate synthase | fps | Synthesis of FDP | Cytosol | AF112881 | Chen et al., 2000; Souret etal., 2002 |
| Sesquitermene cyclases | Catalyzes cyclization of FDP to: | All likely in cytosol | |||
| (i)Epicedrol synthase | eps | 8-Epicedrol | AJ001539 | Mercke et al., 1999; Hua and Matsuda, 1999 | |
| (ii) Amorphadiene synthase | ads | Amorpha-4,11 diene | AJ251751 | Mercke et al., 2000; Chang et al, 2000 | |
| (iii) ?-caryophyllene synthase | cs (qhs1) | ?-Caryophyllene | AF472361 | Cai et al., 2002 | |
| (iv) ?-farnsene synthase | fs | ?-Farnesene | AY835398 | Picaud et al., 2005a | |
| Putative sesquiterpene cyclases | casc125 | Isoprenoid biosynhesis and lyase activity | Isolated from: flowers | AJ271792 | Van geldre, 2000 |
| casc34 | Isoprenoid biosynhesis and lyase activity | leaves and flowers | AJ271793 | Van geldre, 2000 | |
| ses | Reduction product not determined | Young leaves | AAD39832 | Liu et al., 2002 | |
| Squalene synthase | aasqs | Farnesyl-diphosphate farnesyltransferase activity | Endoplasmic reticulum | AY445506 | Liu et al., 2003 |
| Squalene synthase fragment | sqsl | Transferase activity | Cytosol | AF182286 | Souret et al., 2003 |
| CYP71AV1 | na | Catalyzes 3 steps post ADS | Trichomes | DQ315671 | Teoh et al., 2006 |
| PsbA (Fragment) | psba | Act as barcode for flowering plants | Chloroplast | DQ006143 | Kress et al., 2005 |
| Ribulose-1,5-biphosphate Carboxylase/oxygenase | Rbcl | Carbon dioxide fixation; barcoding for flowering plants | Chloroplast | DQ006057 | Kress et al., 2005 |
| Peroxidase 1 | pod1 | Favored the bioconversion of artemisinic acid to artemisinin | Root, stems and leaves | AY208699 | Zhang et al., 2004 |
| Beta-pinene synthase | gh6 | Circadian pattern of expression | Juvenile leaves | AF276072 | Lu et al., 2002 |
| (3R)-linalool synthase | gh1 | Lyase activity | Leaves and flowers | AF154125 | Jia et al., 1999 |
| Isopentenyl transferase | ipt | Biosynthesis of cytokinines phytohormones | Transferred into, A. annua L. via, A. tumefaciens | M91610 | Sa et al., 2001 |
Keeping in view the importance of artemisinin as a novel therapeutic agent for the treatment of drug resistant malaria and its potential to treat other infectious diseases including cancer, we have made an attempt to critically analyze and summarize recent developments related to cellular engineering of artemisinin biosynthetic pathway in this review.
Biosynthetic Pathway of Artemisinin
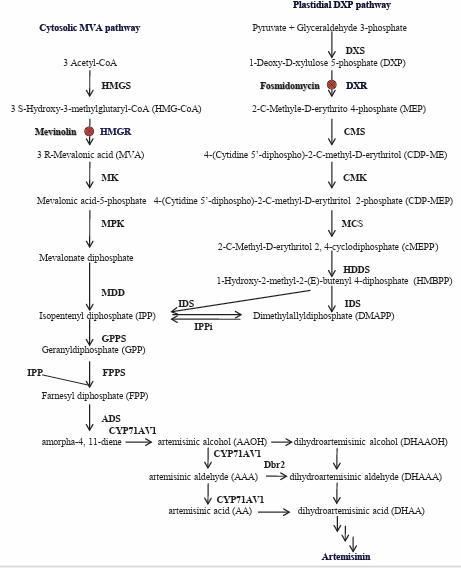
The biosynthetic pathway of artemisinin belongs to the isoprenoid metabolite pathway (Figure 1).
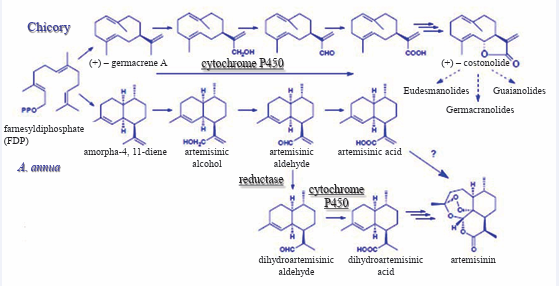
Figure 1: Proposed artemisinin biosynthesis pathway in A. annua L. CMK 4-(Cytidine 5′-diphospho)-2-C-methyl-D-erythritol kinase, CMS 2-C-methyl-D-erythritol 4-phosphate cytidyl transferase, DXR 1-deoxy-D-xylulose 5-phosphate reductoisomerase, DXS 1-deoxy-D-xylulose 5-phosphate synthase, FPPS farnesyl diphosphate synthetase, GPPS geranyl diphosphate synthase, HMGR 3-hydroxy-3-methylglutaryl coenzyme A(HMGCoA) reductase; HMGS HMG-CoA synthase; IDS isopentenyl diphosphate synthase, MCS 2-C-methyl-D-erythritol 2,4-cclodiphosphate synthase, MDD mevalonate diphosphate decarboxylase, MK mevalonate kinase, MPK mevalonate-5-phosphate kinase, SES sesquiterpene synthase, CYP71AV1, cytochrome P450 monooxygenase; Dbr2, artemisinic aldehyde reductase. Adapted from Liu et al., 2006.
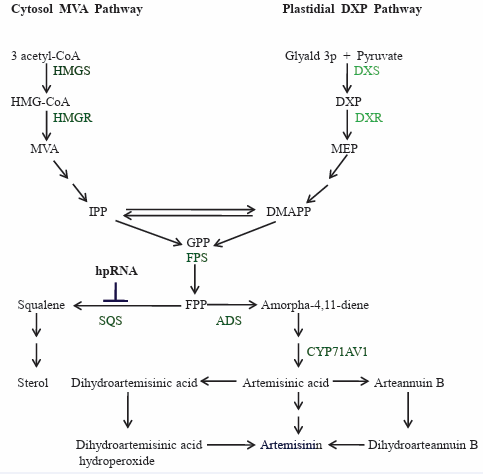
Based on the experimental evidences related to its biosynthesis, artemisinin is suggested to be derived from two common precursors, isopentenyl diphosphate (IPP) and its isomer, dimethylallyl diphosphate (DMAPP). It has been established that higher plants have two independent biosynthetic pathways leading to the formation of IPP: the cytosolic mevalonate pathway and the plastid-localized mevalonate-independent (MEP/Rohmer) pathway (Liu et al., 2005). As a result, mevalonate pathway has no more been considered as the sole route to the synthesis of artemisinin in A. annua L. It was further supported by isolation of two clones encoding deoxy-D-xylulose-5-phosphate synthase (DXPS) and deoxy-D-xylulose-5-phosphate reductoisomerase (DXPR) from transformed hairy roots of A. annua L. [64,97]. The partial carbon supply to the synthesis of artemisinin was reported to be made by MEP pathway operating in plastids and DXR catalyzing the rate limiting step. Recently, the relative contribution of these pathways towards carbon supply in artemisinin production was evaluated by [90]. They demonstrated that mevalonate pathway is the major contributor of carbon and supplies 80% of the carbon to artemisinin biosynthesis, whereas MEP pathway supplies only 20% of the carbon.
In mevalonate pathway, three molecules of acetyl-coenzyme a condensed together to yield 3-hydroxy-3-methylglutaryl CoA (HMG-CoA), which is subsequently, reduced by the enzyme HMG-CoA reductase (HMGR) to yield mevalonic acid (MVA). Then, under the catalysis of mevalonate kinase, mevalonate 5-diphosphate is formed which is subsequently decarboxylated to yield isopentenyl pyrophosphate (IPP) (Newman and Chappell, 1999). The synthesis of IPP and DMAPP by either MVA or DXP pathways is followed by chain elongation. The carbonium ion is a potent alkylating agent that reacts with IPP, giving geranyl diphosphate (GPP). GPP has the active allylic phosphate group and further react with IPP to produce farnesyl pyrophosphate (FPP). FPP takes part in a cyclization reaction catalyzed by cyclases to produce various final products of isoprenoids including artemisinin [27].
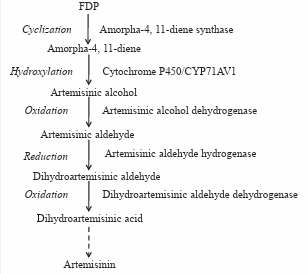
All the steps of mevalonate and MEP pathway have been fully characterized, but post-FPP production of artemisinin is not yet completely elucidated. The formation of the sesquiterpene carbon skeleton, amorpha-4, and 11-diene is catalyzed by amorpha-4, 11-diene synthase [29] for which corresponding cDNAs have been cloned [36,81,108]. The non-descript arrangement of the amorphadiene product belies the unique structural features that ultimately allow for the formation of the 1, 2, 4-trioxane moiety [31] (Figure 2).
Figure 2: Proposed biosynthetic pathway of artemisinin starting from farnesyl diphosphate. On the left is the type of reaction : on the right is the enzyme for each known enzymatic action. Broken arrow indicate multiple steps. Adapted from Bertea et al. (2005).
Expression analysis of CYP71AV1 in A. annua L. tissues indicates that it is most highly expressed in secretory glandular trichomes (GSTs) (Teoh et al., 2006). The moderate expression observed in flower buds presumably reflects their high density of GSTs. Low but detectable levels of RT-PCR products could be observed in leaves. The role of CYP71AV1 in the hydroxylation of amorpha-4, 11-diene is undoubtedly important in artemisinin biosynthesis. The subsequent route to artemisinin is less clear. Most evidence implicates dihydroartemisinic acid as a late precursor to artemisinin biosynthesis, which is derived from artemisinic alcohol by oxidation at C12 and reduction of the C11–C13 double bond. This is based on in vitro biochemical evidence [28], as well as the conversion of dihydroartemisinic acid to artemisinin both in vivo [31] and in vitro in an oxygendependent non-enzymatic fashion.
Bertea and co-workers [28] showed that A. annua L. leaf microsomes convert amorphadiene to artemisinic alcohol in the presence of NADPH. The route from artemisinic alcohol to artemisinin is still not entirely clear, which is evident from the published data reviewed by. In this regard, it is useful to consider the possible route(s) to artemisinin among the pathways shown in Figure 2. These pathways are based on a few conversions whose order may vary. These conversions include the oxidation of C12 from alcohol to aldehyde as well as aldehyde to acid, the reduction of the double bond at C11,13 and the formation of the 1,2,4-trioxane moiety. The later steps in artemisinin biosynthesis remain controversial and theories differ mainly in the identification of either artemisinic acid or dihydroartemisinic acid as the later precursor. The evidence for artemisinic acid has been reviewed by Li et al. (2006). This includes the suggestion that C11, 13 double bond reductions occurs at the level of an intermediate beyond artemisinic acid, such as arteannuin B or artemisitene. On the other hand, the co-occurrence of dihydroartemisinic acid with high artemisinin levels suggests that even if double bond reduction could occur at a very late step, it also occurs in less oxidized precursors. The double bond reduction at C11, 13 is of general interest biochemically, given the relative rarity of enzymes catalyzing double bond reductions. The dihydroartemisinic acid is also being considered as a late precursor of artemisinin biosynthesis. Labelled dihydroartemisinic acid is incorporated into artemisinin in vivo, a sequence which can occur in the absence of enzymes [31,98], 2006; Upstream of dihydroartemisinic acid, the order of oxidations and reduction of artemisinic alcohol en route to dihydroartemisinic acid is still not settled [28] provided biochemical evidence for the fate of artemisinic alcohol in A. annua L. using GST cell-free extracts.
Combinatorial Cellular Engineering of Artemisinin Biosyntheic Pathway
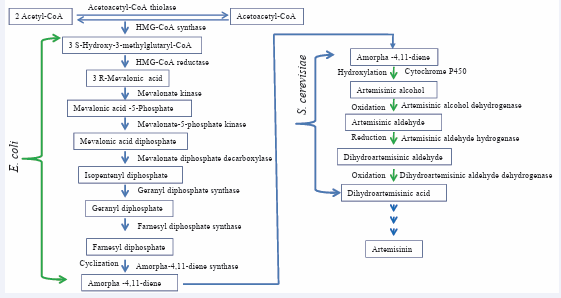
Naturally occurring terpenoids are produced in small quantities, and thus, their purification results in low yields. Further, the complex structures of these molecules make their chemical synthesis challenging and often uneconomical due to poor yields. Transferring metabolic pathways in genetically traceable industrial biological hosts (E. coli and Saccharomyces cerevisiae) offers an attractive alternative to produce large quantities of these complex molecules. To accomplish the production of artemisinin in microbial host altering of MVA and MEP pathways along with addition of very specialized enzymes, e.g. ADS is required. Based on preliminary work on engineering of the MEP pathway to increase isoprenoid precursors for highlevel production of carotenoids [7,18, 51,57,75,] Keasling’s group further developed a base technology for production of amorpha-4, 11-diene in E. coli [78]. Bacterium contains the MEP pathway for production of IPP/DMAPP, but lacks the MVA pathway. Keasling’s group postulated that the MEP pathway is subjected to unknown control elements in bacteria and direct alteration might impair growth. They, therefore, added a truncated MVA pathway from Saccharomyces cerevisiae that was coupled to ADS in E. coli (Figure 3).
Figure 3: Combinatorial biosynthesis of artemisinin starting from acetyl coenzyme A.
It resulted in good bacterial growth and high level production of amorpha-4, 11-diene, estimated up to 100 μg/L in 12 h. Thus these engineered E. coli strains can serve as platform hosts for the production of essentially any terpenoid for which the biosynthetic genes are available, since IPP and DMAPP produced by either arm of the terpenoid pathway are universal precursors to all terpenoids. This strategy has been combined with engineering of genes from the mevalonate dependent isoprenoid pathway (Figure 3), which resulted in an E. coli strain producing 24 μg/ml amorpha-4,11-diene (calculated as caryophyllene equivalent) from acetyl-CoA after supplementation of 0.8% glycerol [78,102] successfully achieved up to 27.4 g/L amorpha-4, 11-diene through E. coli fermentation system. More recently, [101] have isolated the next enzyme in the artemisinin biosynthetic pathway, i.e., cytochrome P450 enzyme (CYP71AV1). This enzyme appears to catalyze the next three steps in artemisinin biosynthesis, an enzymatic function which has also been confirmed by Keasling’s group.
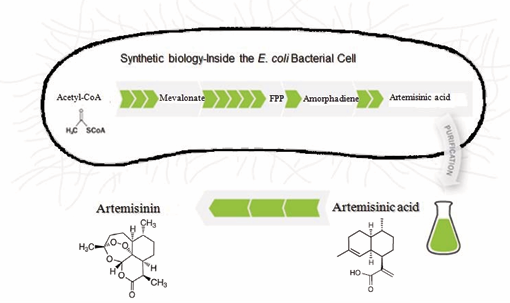
The Bill and Melinda Gates Foundation awarded a five-year grant of $42.6 million in December 2004 to the Institute for One World Health and a non-profit pharmaceutical company (Amyris Biotechnologies), to fund the research and development partnership between Amyris and U.C. Berkeley. The research used synthetic biology to develop a stable and scalable, low-cost technology platform for producing artemisinin and its derivatives. The goal of the collaboration is to create a consistent, high-quality and affordable new source of artemisinin, a key ingredient for making life-saving anti-malarial drugs known as artemisininbased combined therapies (ACTs). In this case, the project team is using synthetic biology to insert genes from the plant A. annua L. into E. coli, a bacterium. Professor Jay Keasling’s laboratory in the Centre for Synthetic Biology at the University of California, Berkeley has completed the synthetic biological process to produce artemisinic acid, a precursor to artemisinin (Figure 4).
Figure 4: Semi-synthetic approach to produce artemisinin and its precursors using E. coli fermentation system.
In another study, attempts have been made to use S. cerevisiae for the production of artemisinin precursors. The expression of the amorpha-4, 11-diene synthase gene in yeast using plasmids and chromosomal integration led to the production of 600 and 100 μg l-1 amorpha-4, 11-diene, respectively after 16 days batch cultivation [66,91] have reported the production of 100 mg/L artemisinic acid in S. cerevisiae strain containing an engineered MVA pathway coupled with the genes encoding amorpha-4, 11-diene synthase and CYP71AV1. This strain transported artemisinic acid, the artemisinin precursor, outside the yeast cell, which makes purification of the product less complex [87] provided major breakthrough using strains of S. cerevisiae (baker’s yeast) and achieved upto 25 gm per litre of artemisinic acid with fermentation and also achieved 40-45% conversion rate of artemisinic acid into artemisinin. Artemisininc acid can be used for the semi-synthesis of artemisinin, but to lower the costs for production of the drug bio processing must be optimized [70].
Genetic engineering of A. annua L. plant
It is envisaged to produce high artemisinin yielding transgenic strains of A. annua L. plants, which will ensure a constant high production of artemisinin by over-expressing the key enzymes in the terpene and artemisinin biosynthetic pathways, or by inhibiting enzyme(s) of another pathway competing for artemisinin precursors. In recent years, remarkable progress has been made in the understanding of molecular biology of artemisinin biosynthesis and its regulation [29,103]. The genes of the key enzymes involved in the biosynthesis of artemisinin, such as HMG-CoA reductase, farnesyl pyrophosphate synthase (FPS), amorpha-4, 11-diene synthase (ADS), and the genes of the enzymes involved in the pathway competing for artemisinin precursors, such as squalene synthase (SQS) involved in sterol biosynthesis, have been cloned from A. annua L. [18,73,80,81,108]. On the other hand, [89,111] induced hairy roots in A. annua L. employing Agrobacterium rhizogenes. Further, the factors influencing transformation efficiency of A. rhizogenes were explored to optimize the transformation system by [71,117] induced hairy root in A. annua L. leaf blade pieces and petiole segments infected with A. rhizogenes strain 1601 and obtained a clone with high content of artemisinin (1.195 mg/g DW).
To develop transgenic A. annua L. strains with high content of artemisinin by modulating the expression of above mentioned genes, an efficient system of genetic transformation as well as regeneration of ex plants of A. annua L. should be in place [105] developed an A. tumefaciens-mediated transformation system for A. annua L. plants with high transformation rates (75% regenerants harboring foreign gene). Artemisinin content in the leaves of regenerated plants was 0.17%, a little bit higher than that present in the leaves of normally cultured plants (0.11% DW). They further investigated the factors viz., the age of explants, A. tumefaciens strain and plant genotype influencing the transformation efficiency [106]. Later, [55] established a high efficiency genetic transformation and regeneration system for A. annua L. via A. tumefaciens.
Artemisinic acid is one of the precursors of biosynthesis of artemisinin, which has the cadinene structure. [39] transformed a cotton cadinene synthase cDNA into the leaf explants of A. annua L. using A. rhizogenes. In the isoprenoid biosynthesis pathway, farnesyl pyrophosphate synthase (FPS) catalyzes the two sequential 1-4 condensations of IPP with DMAPP to produce GPP and with GPP to give FPP, which is then utilized by isoprenoid pathway and artemisinin biosynthetic pathway to produce isoprenoids and artemisinin, respectively [34]. The cDNAs encoding FPS have been isolated from a number of plant species, including Arabidopsis thaliana [43] and Lupinus albus [25]. Since 15-carbon FPP can be catalyzed by sesquiterpene cyclises, such as, ADS to form cyclic sesquiterpenoids (amorpha-4, 11-diene in A. annua L.), overexpressing foreign FPS gene into A. annua L. plants holds the possibility of affecting accumulation of artemisinin. A cDNA encoding cotton FPPS placed under a CaMV 35S promoter was hence, transferred into A. annua L. Plants via A. tumefaciens strain LBA 4404 or A. rhizogenes strain ATCC 15834 mediated genetic transformation [37,38]. In the transgenic plants, the concentration of artemisinin was approximately 8-10 mg/g DW, which were 2 to 3-folds higher than that in the control plants. [56] achieved about 34.4% increase in artemisinin content by over expressing FPS. We have over expressed one of the key regulatory enzyme of MVA pathway (HMGR) in A. annua L. plants via A. tumefaciens mediated transformation and achieved 39% enhancement in artemisinin contents as compared to control plants [100,59] simultaneously over expressed cyp71av1 and cpr genes in A. annua L. and recorded 2.4 fold enhancement in artemisinin content. The cytokinin biosynthetic gene codes for the enzyme isopentenyl transferase (ipt), which catalyzes the condensation of isopentenyl pyrophosphate and adenosine monophosphate (AMP) to yield isopentenyl AMP is believed to represent the rate-limiting step in cytokinin biosynthesis in tumorous plant tissue [21,22]. The influence of over expression of isopentenyl transferase gene on the physiological and biochemical characteristics of A. annua L. plant was studied by. The transgenic A. annua L. plants were found to accumulate more cytokinins (2 to 3-fold), chlorophyll (20-60%) and artemisinin (30–70 %), when compared with control plants. Previous studies indicated that capitate glands on the leaf surface [49] and specialized chloroplasts of the capitate gland appeared to play very important role in artemisinin biosynthesis [48]. Light affects to terpene biosynthesis in general and artemisinin biosynthesis in particular by modulating carbon flux through regulation of HMGCoA reductase, a key regulatory enzyme in mevalonate pathway. In case of potato, it has been reported that light regulates HMGR at both transcriptional and translational level. In A. annua L., β-pinene synthase was found to have a circadian pattern of gene expression, accompanied by a similar temporal pattern of β-pinene emission under light exerting a stimulatory effect. Analysis of root cultures of A. annua L. suggested that light also positively regulates artemisinin biosynthesis because the root cultures exhibited a substantial decrease in artemisinin content when moved from light to dark [69]. [58] hence, over expressed Arabidopsis blue light receptor CRY1 in A. annua L. to evaluate its effect on artemisinin synthesis and accumulation. They found that over expression of CYP1 gene had resulted in increased accumulation of both artemisinin (30-40%) and anthocyanins (2- fold) as compared to control plants. [116] co-over expressed dxr, cyp71av1 and cpr in A. annua L. plant and achieved three time higher level of artemisinin in compare to wild type A. annua line.
Genetic Engineering of Cychorium intybus for Artemisinin Production
Dafra Pharma International NV and Plant Research International (PRI) have initiated new research to produce artemisinin via genetically modified chicory plants. In studies carried out at Wageningen, the complete biosynthetic pathway of artemisinin was resolved [42,28] (Figure 5).
Figure 5: Biosynthetic routes of bitter sesquiterpene lactones in chicory and artemisinin in Artemisia annua L. adapted from Bertea et al. (2005) and de Kraker et al. (2003).
In addition, the Wageningen group, headed by Prof. Harro Bouwmeester and Dr. Maurice Franssen, demonstrated that chicory enzyme(s) normally involved in the biosynthesis of the bitter sesquiterpene lactones in chicory, were capable of performing reactions required for the biosynthesis of artemisinin [42]. The group of Prof. Bouwmeester has tried to produce the chemical precursor for artemisinin (dihydroartemisinic acid) in the roots of chicory via a diversion of the biosynthesis of bitter compounds. On the other hand, the group of Prof. Bouwmeester has shown in a wide range of plant species that diversion of the biosynthesis of terpenes can be carried out very efficiently [62]. Moreover, they also demonstrated that upto 40 kg ha-1 dihydroartemisinic acid can be produced using genetically modified chicory.
RNAi (RNA interference) Mediated Gene Silencing
The mechanism of artemisinin biosynthesis has recently become much clearer [18,28,68]. It has been shown that artemisinin belongs to the isoprenoid group of compounds, which are derived from two common precursors, namely, IPP (isopentenyl diphosphate) and its isomer DMAPP (dimethylallyl diphosphate). GPP (geranyl diphosphate) is formed by chain elongation from IPP and DMAPP when they react with a carbonium ion, and GPP can then further react with IPP to produce FPP (farnesyl diphosphate). FPP can be converted through enzymic catalysis to produce various isoprenoid final products, such as, artemisinin and sterols. SQS (squalene synthase) is reported to be the key enzyme catalyzing first step of the sterol biosynthetic pathway, a pathway competing with artemisinin biosynthesis (Figure 6).
Figure 6: RNAi mediated suppression of SQS gene. Adapted from Zhang et al. 2009.
The gene for SQS has been cloned from A. annua L. [73]. Previous studies have shown that inhibition of the sterol biosynthetic pathway by chemical methods could improve artemisinin content of A. annua L. [65,114] showed in their study that artemisinin production could be enhanced by the addition of naphtiphine, an inhibitor of the enzyme squalene epoxidase, to the medium.[65] also demonstrated that the application of many sterol inhibitors, including miconazole or chlorocholine, resulted in an increase in artemisinin in shoot cultures of A. annua L. Recently, [91] confirmed that downregulation of ERG9 (ergosterol biosynthesis-pathway gene 9), a gene that encodes SQS in yeast, using a methionine-repressible promoter (PMET3), increased the production of amorpha-4,11- diene 2-fold in a yeast strain into which FPP-synthetic-pathway genes and the ADS (amorpha-4,11-diene synthase) gene from A. annua L. had been incorporated. RNAi mediated by hpRNA has been used in gene silencing in many species of plants [69] reported that hpRNA mediated down-regulation of ghSAD-1 and ghFAD2-1, two key enzymes in the fatty-acid-biosynthesis pathway in cotton (Gossypium hirsutum), elevated the stearic acid content (44% compared to a normal level of 2%) and oleic acid content (77% compared with a normal level of 15%) in cotton seeds. It was also reported that suppression of one key enzyme, CaMXMT1, involved in the caffeine-biosynthetic pathway through hpRNA-mediated interference in coffee (Coffea spp.) decreased obromine and caffeine accumulation, efficiently 30–50% of that normally found in the species [85]. In a study with opium poppy (Papaver somniferum), an hpRNA construct containing sequences from multiple cDNAs of genes in the codeine reductase gene family was used to silence several enzymes in the pathway. In the developed transgenic plants, the non-narcotic alkaloid (S)-reticuline, which occurs upstream of codeine in the pathway, accumulated at the expense of morphine, codeine, opium and thebaine [23]. In tomato (Solanum lycopersicum), hp (hairpin) construct was used to suppress an endogenous photo morphogenesis regulatory gene, DET1, driven by a fruit-specific promoter. DET1 was degraded and the carotenoid and flavonoid content of tomato fruits were increased, while all other traits for fruit quality remained unchanged in transgenic plants compared with that in wild-type tomato [41]. In another study, suppression of an arsenic reductase gene ACR2 in Arabidopsis (thale cress) using hp constructs improved the arsenic content significantly in transgenic shoots (10–16-fold compared with that in the wildtype) [45]. Following the similar strategy, artemisinin content was enhanced upto 2-3 fold in transgenic A. annua L. plants (≈ 31.4 mg g-1dw as compared to 8-10 mg g-1dw in control plants) by suppressing the expression of sqs (squalene synthase gene), encoding SQS, a key enzyme of sterol pathway (a pathway competitive with that of artemisinin biosynthesis) by means of hp-RNA-mediated RNAi (RNA interference) [120] (Figure 6). The sterol content of transgenic plants was also reduced to 37 - 58% as compared to the wild type plants, but it had not affected their growth and development. This study along with others, therefore, demonstrates that the metabolic engineering strategy of suppressing sterol biosynthesis using RNAi could become an effective and suitable mean for increasing the artemisinin content of plants.
Role of Transcription Factors in Artemisinin Biosynthesis
The functions of an increasing number of plant transcription factors are being elucidated, and many of these factors have been found to impact flux through metabolic pathways. Since transcription factors, as opposed to most structural genes, tend to control multiple steps pathway therefore, they have emerged as powerful tools for the manipulation of complex metabolic pathways in plants. The importance of transcription factors in the regulation of the flavonoid pathway suggests that they may play an equally important role in regulating other pathways of plant secondary metabolism [32]. In several species, relevant observation is the fact that terpenoid accumulation is preceded by the coordinated induction of several pathway genes. It is likely that several aspects of terpenoid metabolism are regulated at the level of gene expression, while relatively little is known of the transcription factors being involved. The first direct evidence of transcription factor control over terpenoid pathway gene expression was observed in Catharanthus cells over-expressing (mono) terpenoid indole alkaloid (TIA) pathway activator ORCA3 [104]. Analysis of transcript levels showed that, in addition to TIA pathway genes, the gene encoding DXS was also induced. Although induction was found to be significant, it was also fairly limited. G10H encoding geraniol 10-hydroxylase, another gene in the monoterpene branch of the pathway, which was monitored in this experiment, was not affected. These observations suggest a role for ORCA3 in regulating terpenoid as well as TIA biosynthesis, although additional factors are likely to be involved [104]. Recently, [76] isolated and characterized AaWRKY1, an A. annua L. transcription factor that regulated the amorpha-4, 11-diene synthase gene, a key gene of artemisinin biosynthesis. Promoters of ADS contain two reverse-oriented TTGACC W-box cis-acting elements, which are binding sites of WRKY transcription factors. A full length cDNA (AaWRKY1) was isolated from a cDNA library of the glandular secretory trichomes (GSTs) in which artemisinin is synthesized and sequenstered. AaWRKY1 and ADS genes were highly expressed in GSTs and both were strongly induced by methyl jasmonate and chitosan. [76] also demonstrated that AaWRKY1 has a similar propensity. Transient expression of AaWRKY1 activated the expression of HMGR, ADS, CYP71AV1 and DBR2 of the artemisinin biosynthesis pathway. It is possible that the W-box also existed in the promoters of CYP71AV1, HMGR and DBR2 in A. annua L. Indeed, the W-box has been found in the promoter of cytochrome P450 genes of many other plants such as Arabidopsis [83], C. japonica [63] and cotton [118], while two W-box elements were also found in the HMGR1 promoter of Camptotheca acuminate [33].
CONCLUSIONS
A. annua L. is the main source of artemisinin which is the most potent and efficacious antimalarial drug after quinine. Artemisinin has also been demonstrated as a selective anticancer drug. Currently, the limited availability of artemisinin and the lack of real competition among producers of raw material seem to be the major barriers to scaling-up production and are partially responsible for its high price [115]. The relatively low yield of artemisinin in A. annua L. and non-availability of an economically viable synthetic protocol have been the major obstacles for its commercial production and clinical use. However, multipoint cellular engineering is now beginning to supersede single-point engineering as the best way to manipulate metabolic flux to enhance the artemisinin synthesis in transgenic plants. It is evident from the foregoing discussion that, several points in a given metabolic pathway can be controlled simultaneously either by over expressing and/or suppressing several enzymes or through the use of transcriptional regulators to control several endogenous genes. Moreover, applied genomics, proteomics and metabolomics are continuing to expand our knowledge of metabolic pathways, while advances in systems biology help us to model the impact of different modifications made at the gene level more accurately
FUTURE PROSPECTS
Cellular engineering of biosynthetic pathway of artemisinin has shown promising results. Because of the intricate and highly complicated metabolic networks of artemisinin pathway in A. annua L., there exists multiple rate-limiting steps in the pathway. The fluxes through the pathway are controlled to a great extent at the level of genes, enzymes, compartmentation, transport and accumulation [107]. It has been established that the enzyme(s) HMGR, IPT, FPS, ADS, CYP71AV1,DXR and CPR are catalyzing for the rate-limiting steps in isoprenoid and artemisinin biosynthesis [24,53,26,56,74,116]. The over-expression of genes for these enzymes in transgenic A. annua L. plants has resulted in considerable increase in artemisinin content. The down regulation of sterol biosynthesis by suppressing genes encoding key enzymes, such as, SQS, has also been shown to enhance artemisinin content. These cellular engineering steps, when used together, could be able to enhance artemisinin content in A. annua L. The other way to enhance artemisinin biosynthesis in A. annua L. could be the use of transcription factors (AaWRKY1 and ORCA3), which are involved in up-regulation of artemisinin biosynthetic pathway and down regulation of sterol biosynthesis. Isolation and characterization of the promoters of HMGR, CYP71AV1, DBR2 and other genes in the artemisinin biosynthetic pathway would be beneficial for further evaluation of the function of WRKY transcription factors in artemisinin metabolism.
Moreover, combinatorial biosynthesis has been utilized for the production of important classes of natural products, including alkaloids (vinblastine and vincristine), terpenoids (artemisinin and paclitaxel) and flavanoids. It is expected, therefore, that combinatorial biosynthetic strategies will yield interesting alternatives in the near future.
The World Health Organization recommends artemisininbased combination therapies (ACTs) as first-line treatment for malaria. However, supplies of plant-derived artemisinin are subject to the seasonality and volatility common to many plant-based commodities, leading to fluctuations in the price of artemisinin. Commercial scale production of semi-synthetic artemisinin would have the potential to stabilize supply and supplement existing plant-derived materials to create a consistent, high-quality and affordable new source of artemisinin and can help to meet the projected world-wide demand for ACTs. Although progress has been made in metabolic engineering of artemisinin biosynthesis in A. annua L., more needs to be done in order to further increase artemisinin production to a practical application through metabolic regulation.
ACKNOWLEDGMENTS
The Research Fellowships to Mr. Mauji Ram by Council of Scientific and Industrial Research, India and Department of Biotechnology, Ministry of Science & Technology, Government of India are gratefully acknowledged. The authors are also thankful to Dr. M.A.A. Khan, NISCAIR, and New Delhi, India for editing the manuscript.
REFERENCES
27. Barkovich R, Liao JC. Metabolic engineering of isoprenoids. Metab Eng. 2001; 3: 27-39.
34. Cane D.E. Enzymatic formation of sesquiterpenes. Chem. Rev. 1990; 90: 1089–1103.
44. DellaPenna D. Plant metabolic engineering. Plant Physiol. 2001; 125: 160-163.
47. Dixon RA. Engineering of plant natural product pathways. Curr Opin Plant Biol. 2005; 8: 329-336.
82. Nair MS, Basile DV. Bioconversion of arteannuin B to artemisinin. J Nat Prod. 1993; 56: 1559-1566.